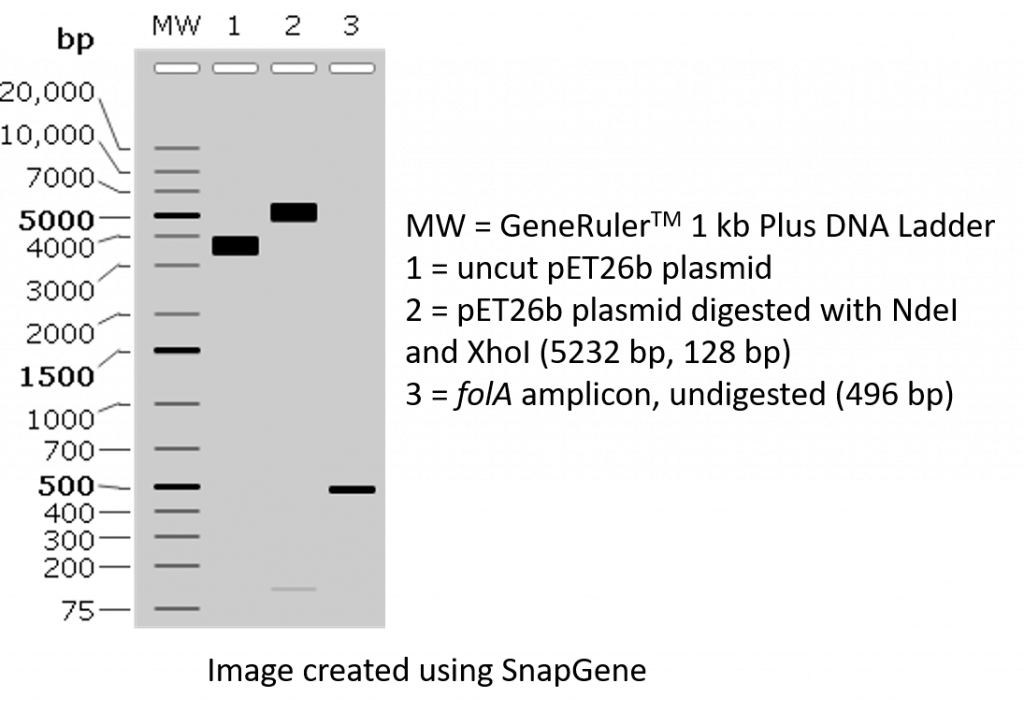
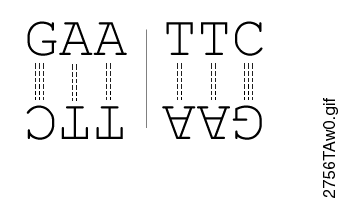
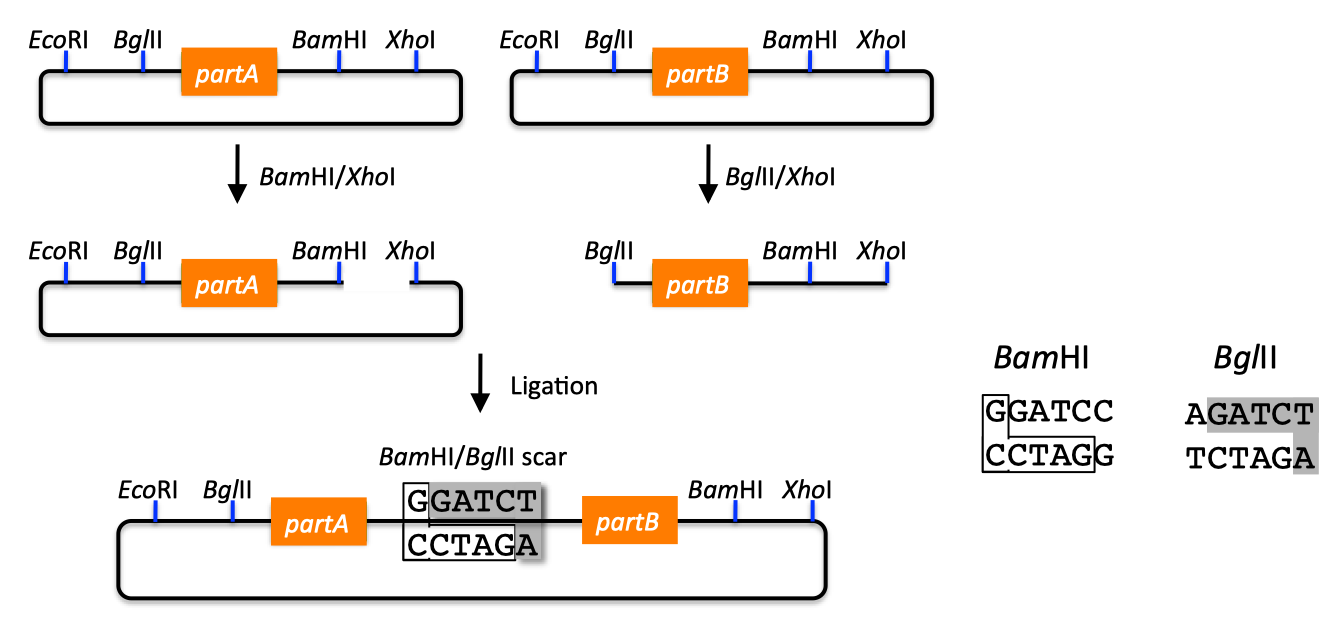
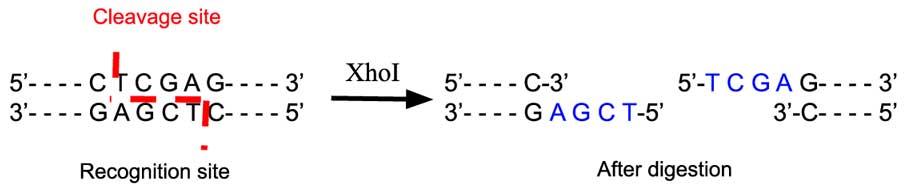
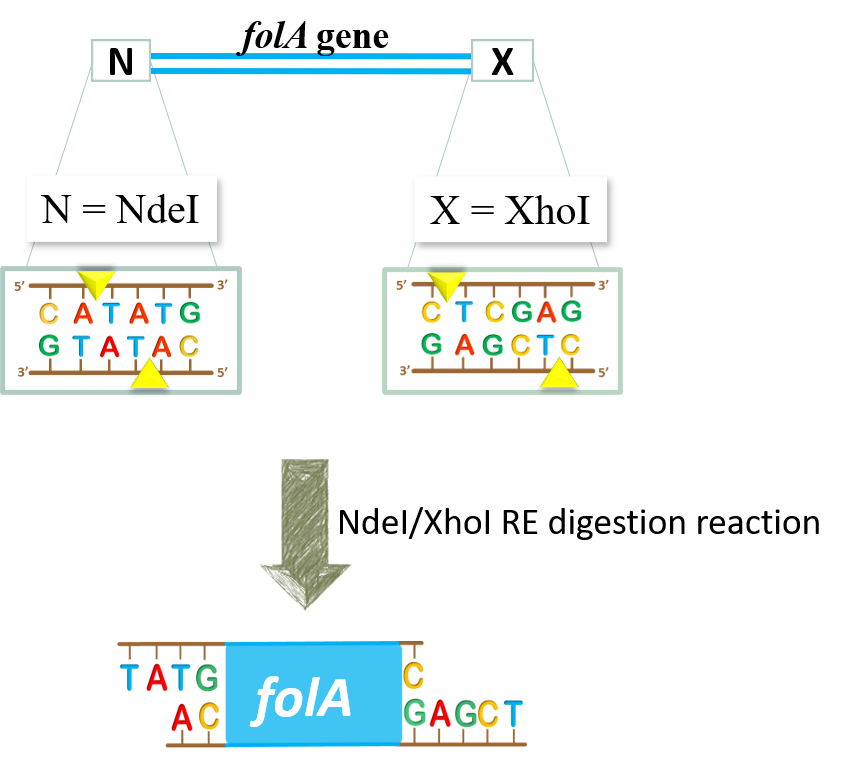
The restriction enzymes Xho I and Xba I were used to digest a linear fragment of DNA at specific sequences such as that shown for EcoRI (figure above).Use the information gained from
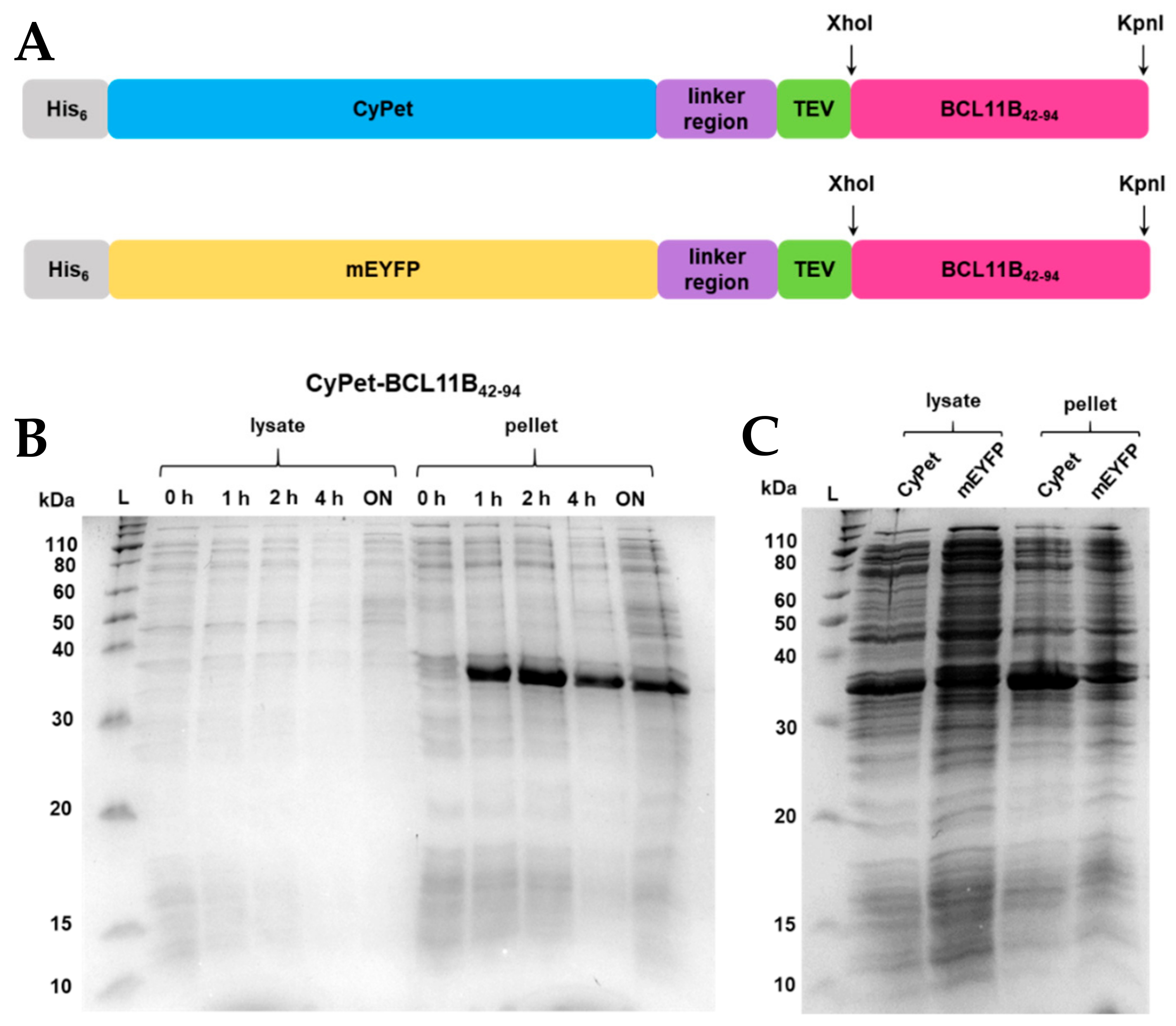
Anne's paper on Easy Expression and Purification of Fluorescent N-Terminal BCL11B CCHC Zinc Finger Domain is published - Fakultät - Universität Greifswald

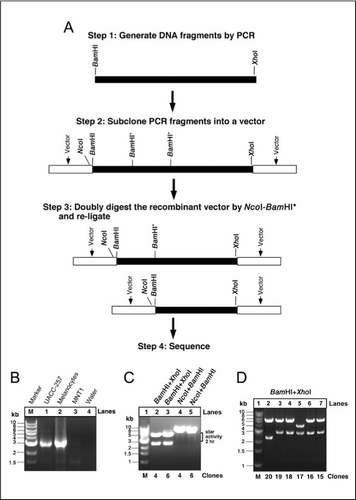
SciELO - Brasil - In-Situ Gel-Free Plasmid Reassembling for Rapid Gene Subcloning and Truncation In-Situ Gel-Free Plasmid Reassembling for Rapid Gene Subcloning and Truncation

SciELO - Brasil - In-Situ Gel-Free Plasmid Reassembling for Rapid Gene Subcloning and Truncation In-Situ Gel-Free Plasmid Reassembling for Rapid Gene Subcloning and Truncation
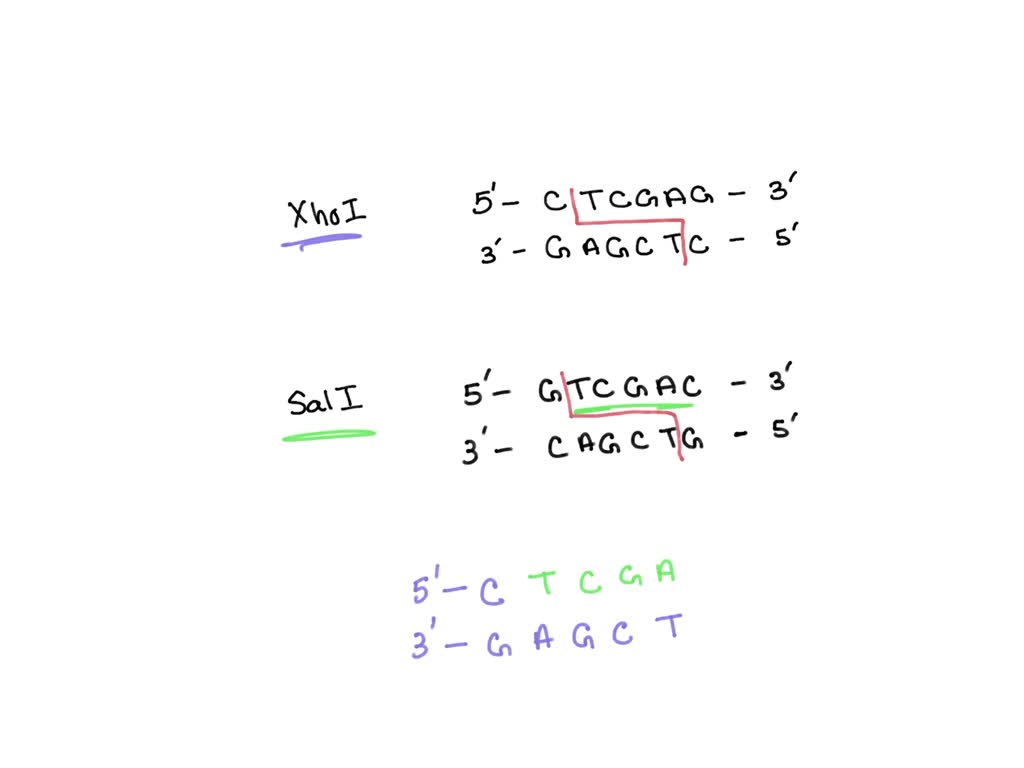
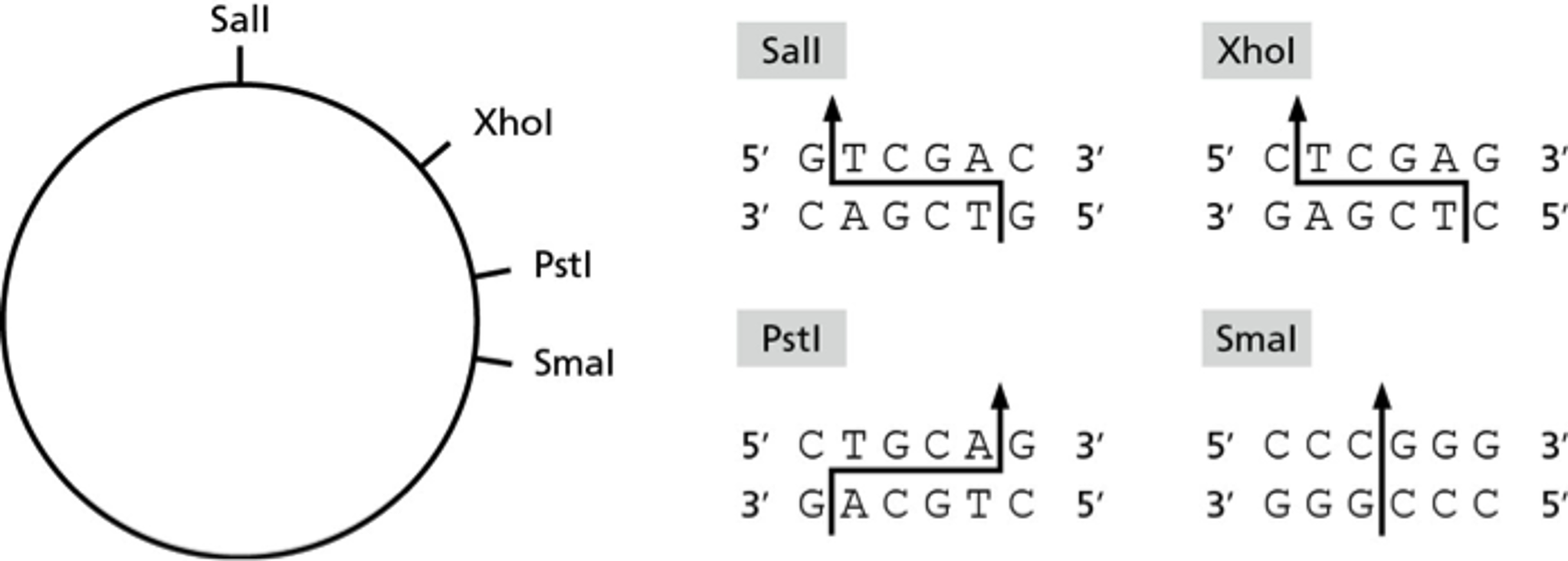
SOLVED: #16) The restriction enzymes Xhol and SalI cut their specific sequences as shown below: XhoI 5' C | TCGAG 3' SalI | 5' GTCGAC 3' 3' GAGC | TC 5' 3'
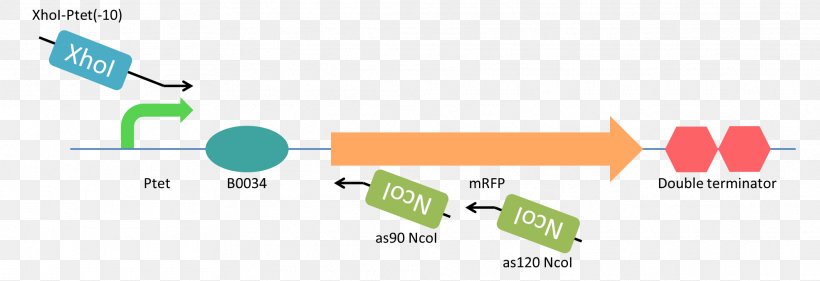
XhoI Restriction Enzyme Antisense RNA Restriction Site Nucleic Acid, PNG, 1972x677px, Restriction Enzyme, Antisense Rna, Area,

Primer sequences. Cleavage sites for the restriction enzymes NheI and... | Download Scientific Diagram

A) Restriction enzyme digestion with Xho I and Nco I of the constructs... | Download Scientific Diagram