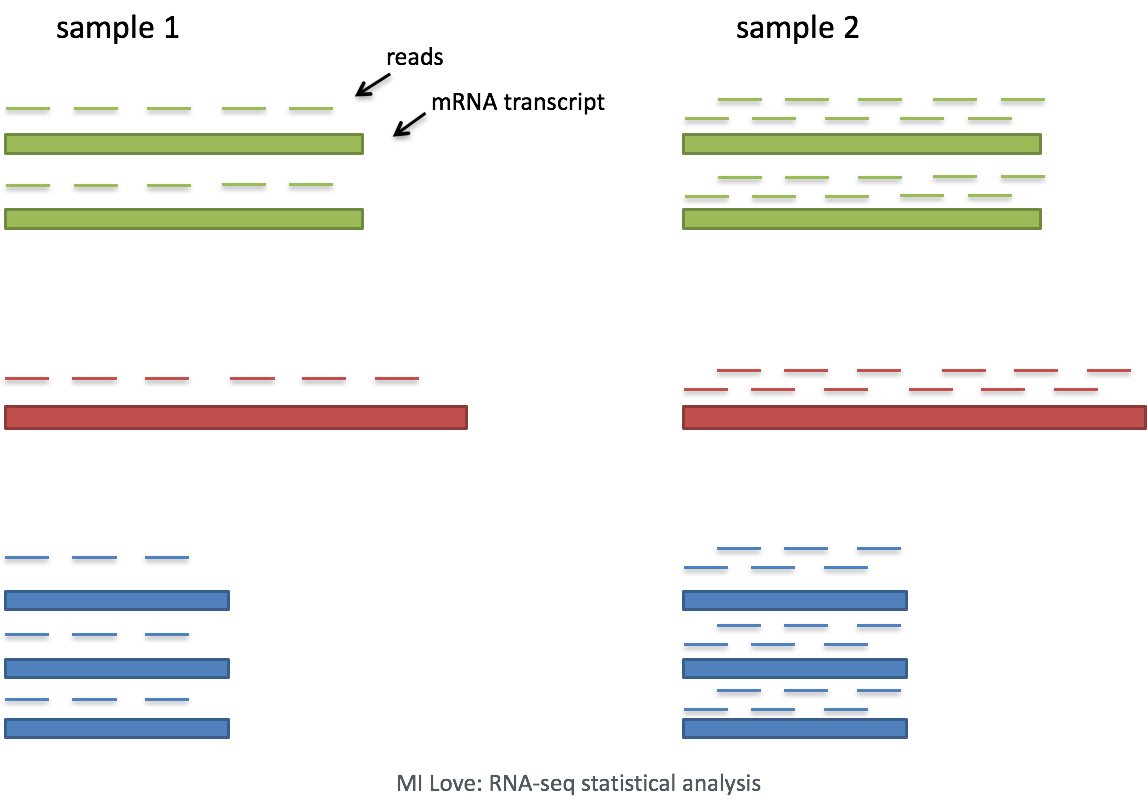
Single-cell RNA-seq: Normalization, identification of most variable genes | Introduction to single-cell RNA-seq

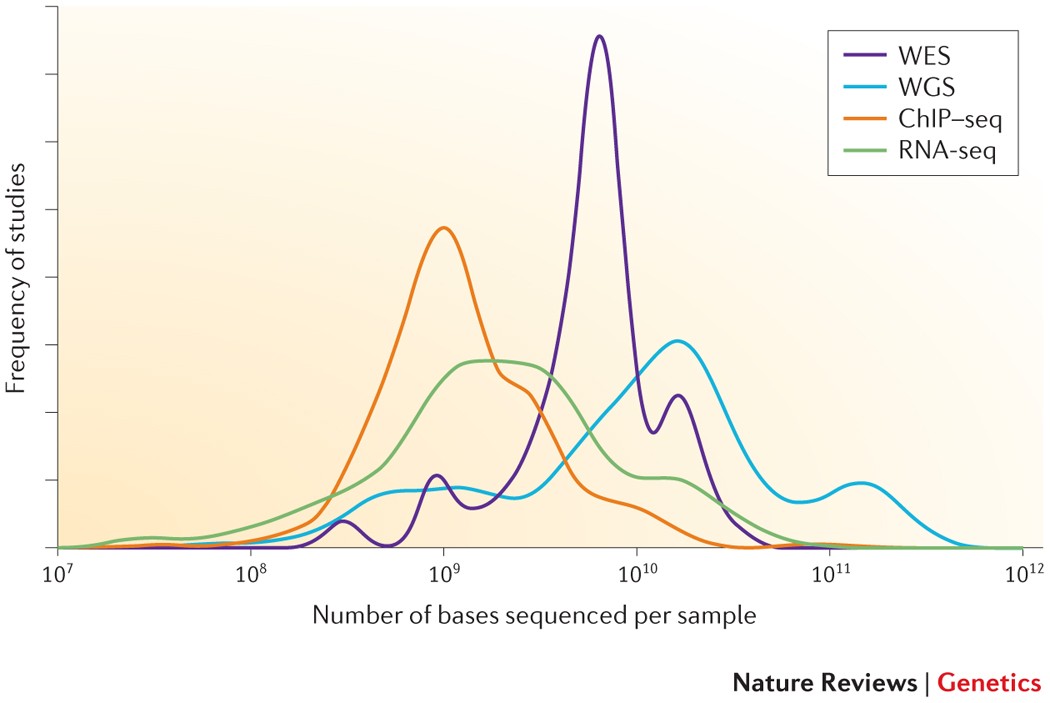
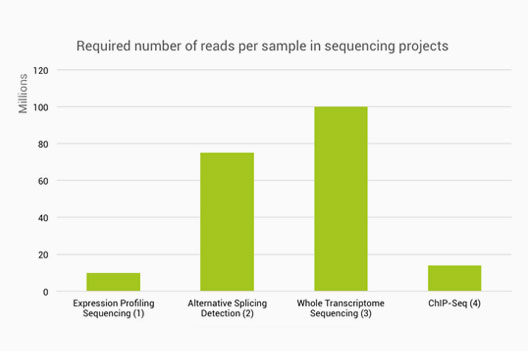
ScienceVision - Different RNA-Seq experiment types require different sequencing read lengths and depth (number of reads per sample). This bulletin reviews RNA sequencing considerations and offers resources for planning RNA-Seq experiments. Link:
Boxplot of sequencing depth data and amplicons size (bp). The range of... | Download Scientific Diagram
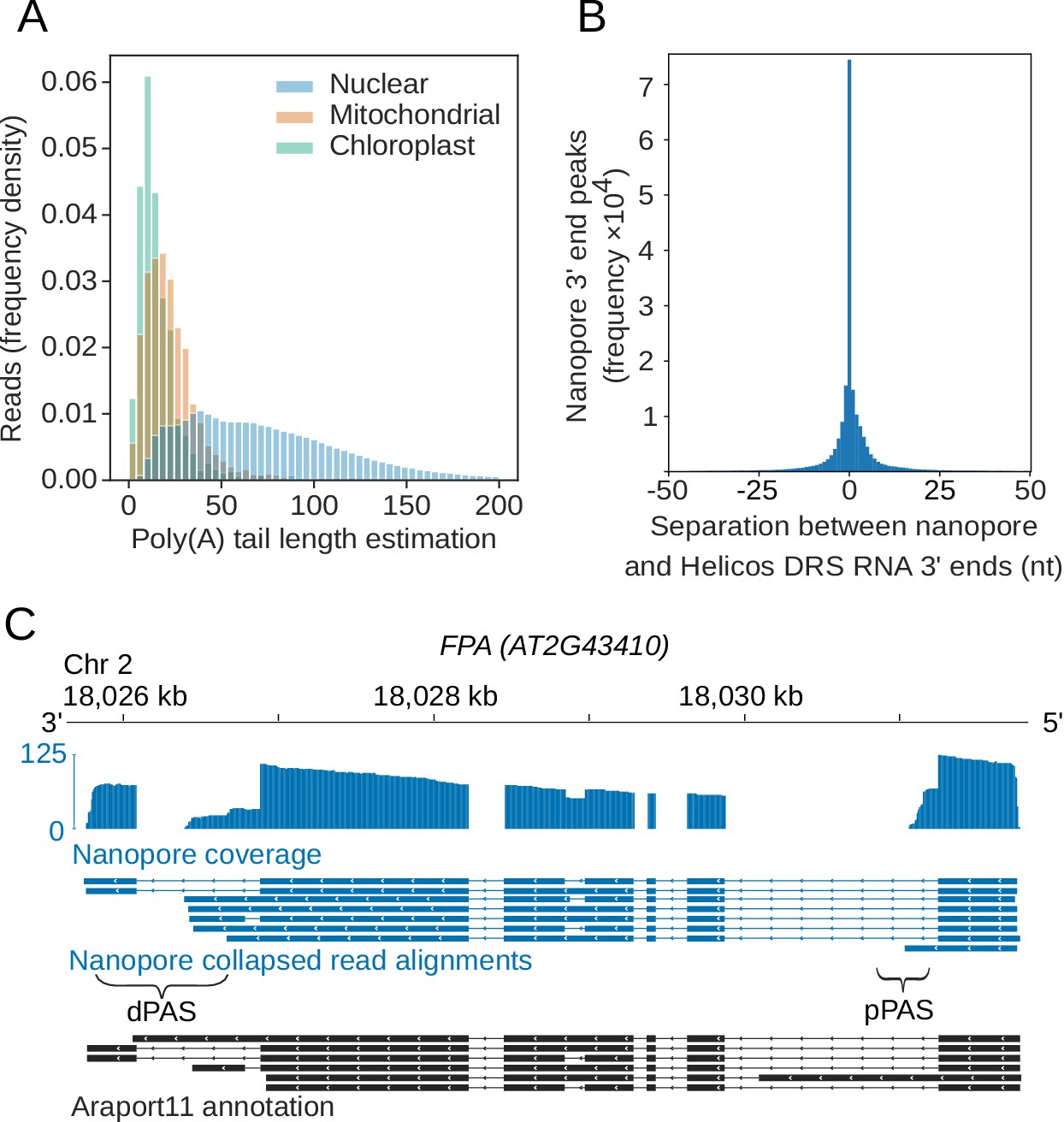
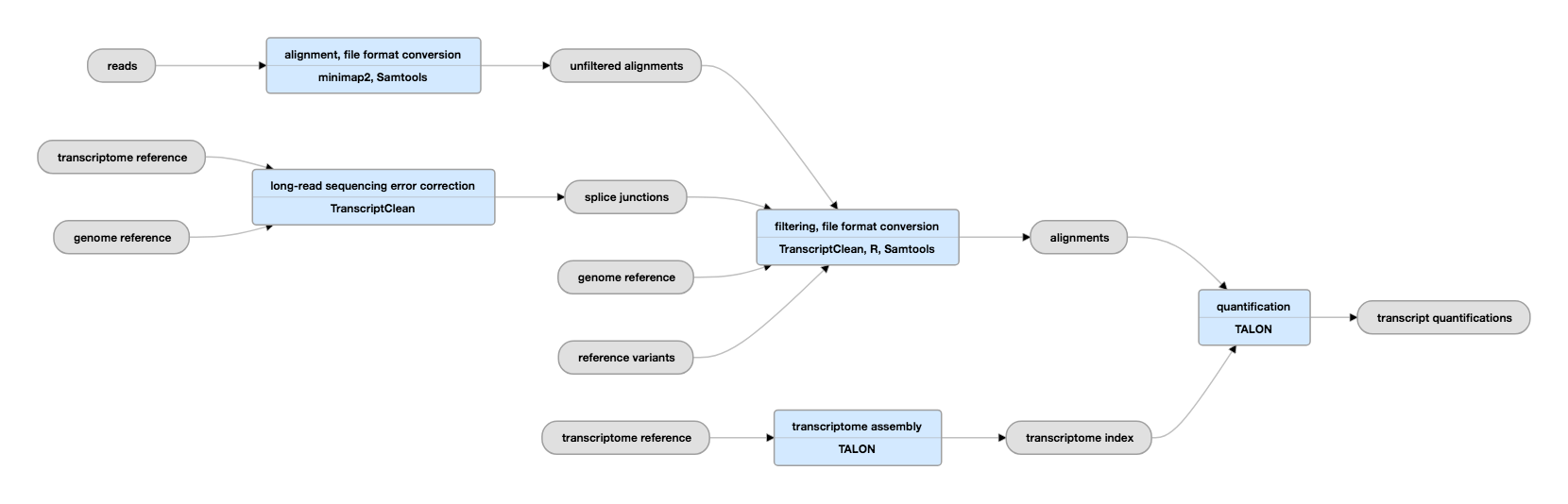
Nanopore direct RNA sequencing maps the complexity of Arabidopsis mRNA processing and m6A modification | eLife

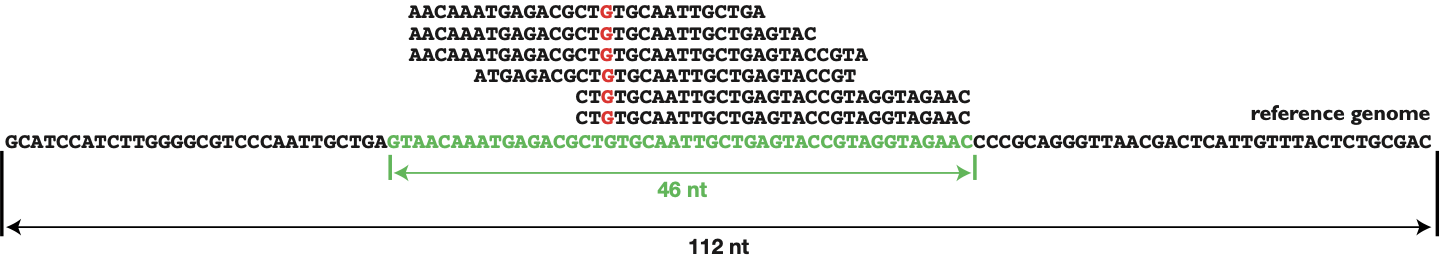
Recommendations for accurate genotyping of SARS-CoV-2 using amplicon-based sequencing of clinical samples - ScienceDirect