IJMS | Free Full-Text | Improved Binding Affinity of Omicron’s Spike Protein for the Human Angiotensin-Converting Enzyme 2 Receptor Is the Key behind Its Increased Virulence
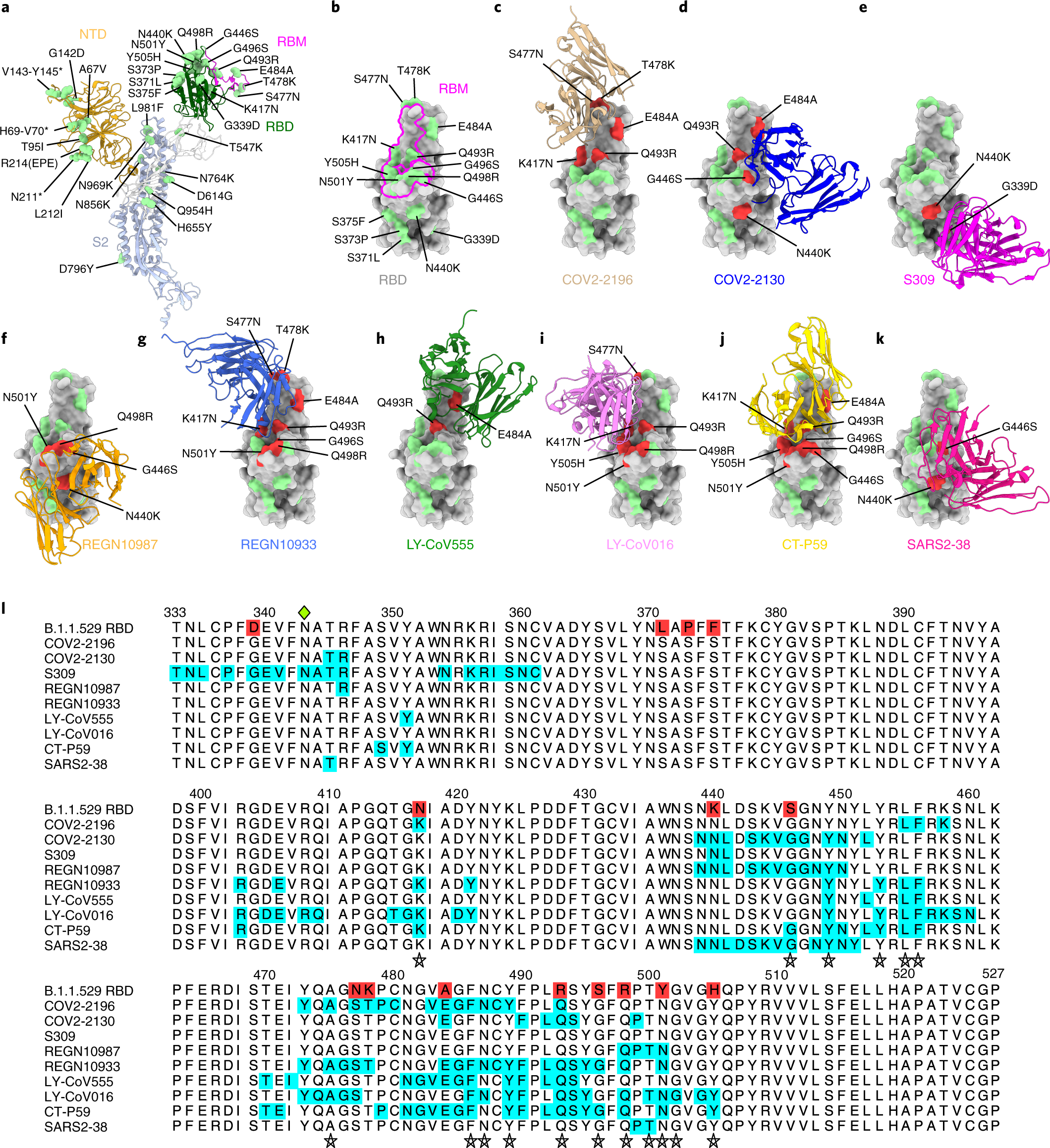

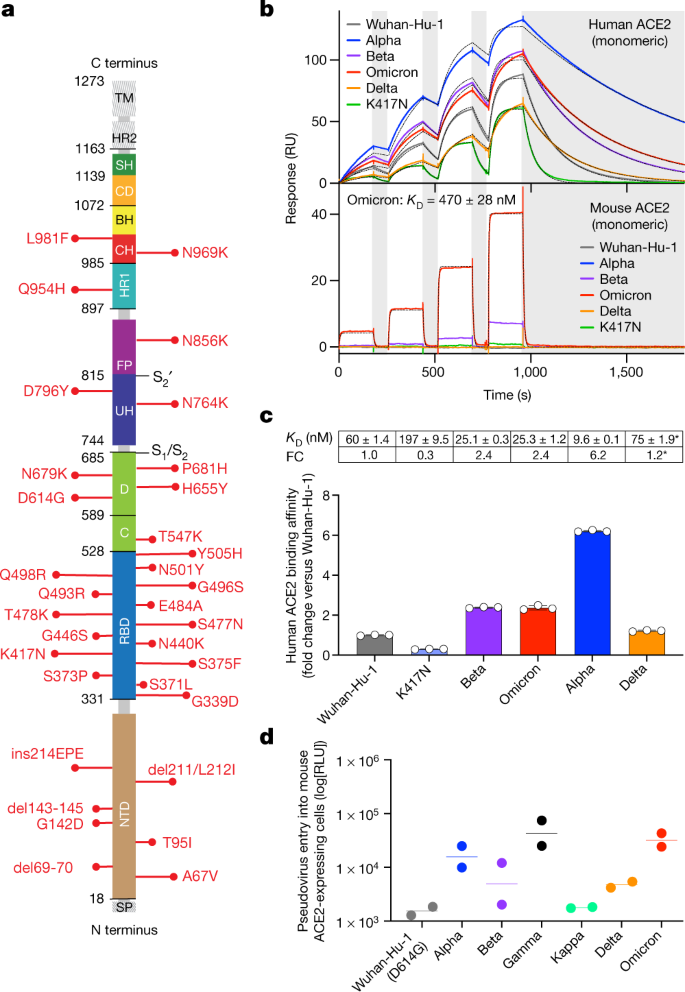
An infectious SARS-CoV-2 B.1.1.529 Omicron virus escapes neutralization by therapeutic monoclonal antibodies | Nature Medicine

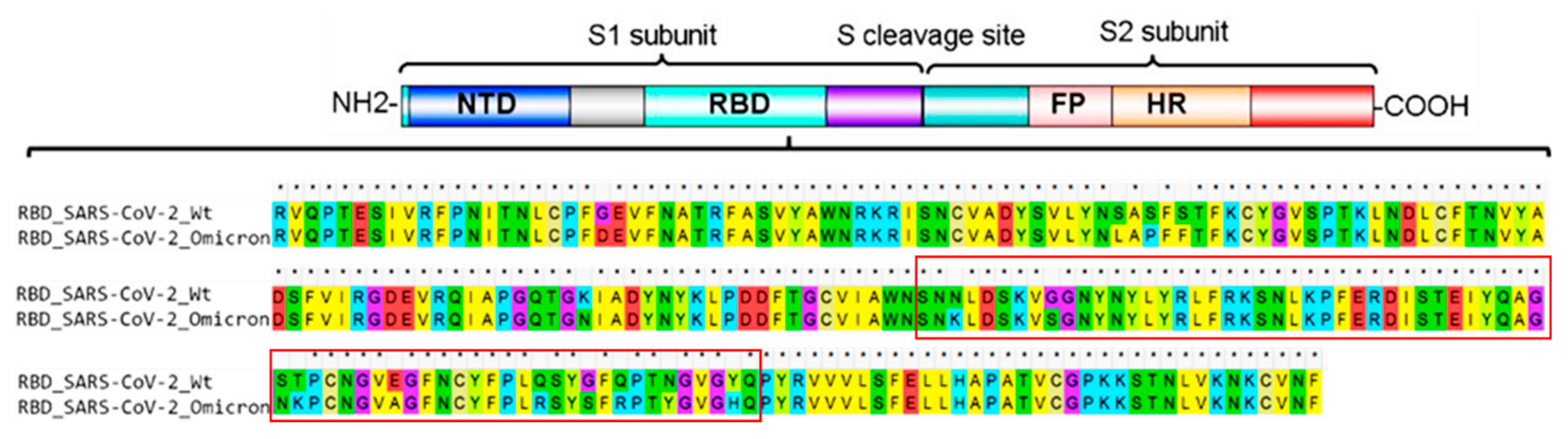
Antigenicity comparison of SARS‐CoV‐2 Omicron sublineages with other variants contained multiple mutations in RBD - Li - 2022 - MedComm - Wiley Online Library

Receptor binding and complex structures of human ACE2 to spike RBD from omicron and delta SARS-CoV-2 - ScienceDirect
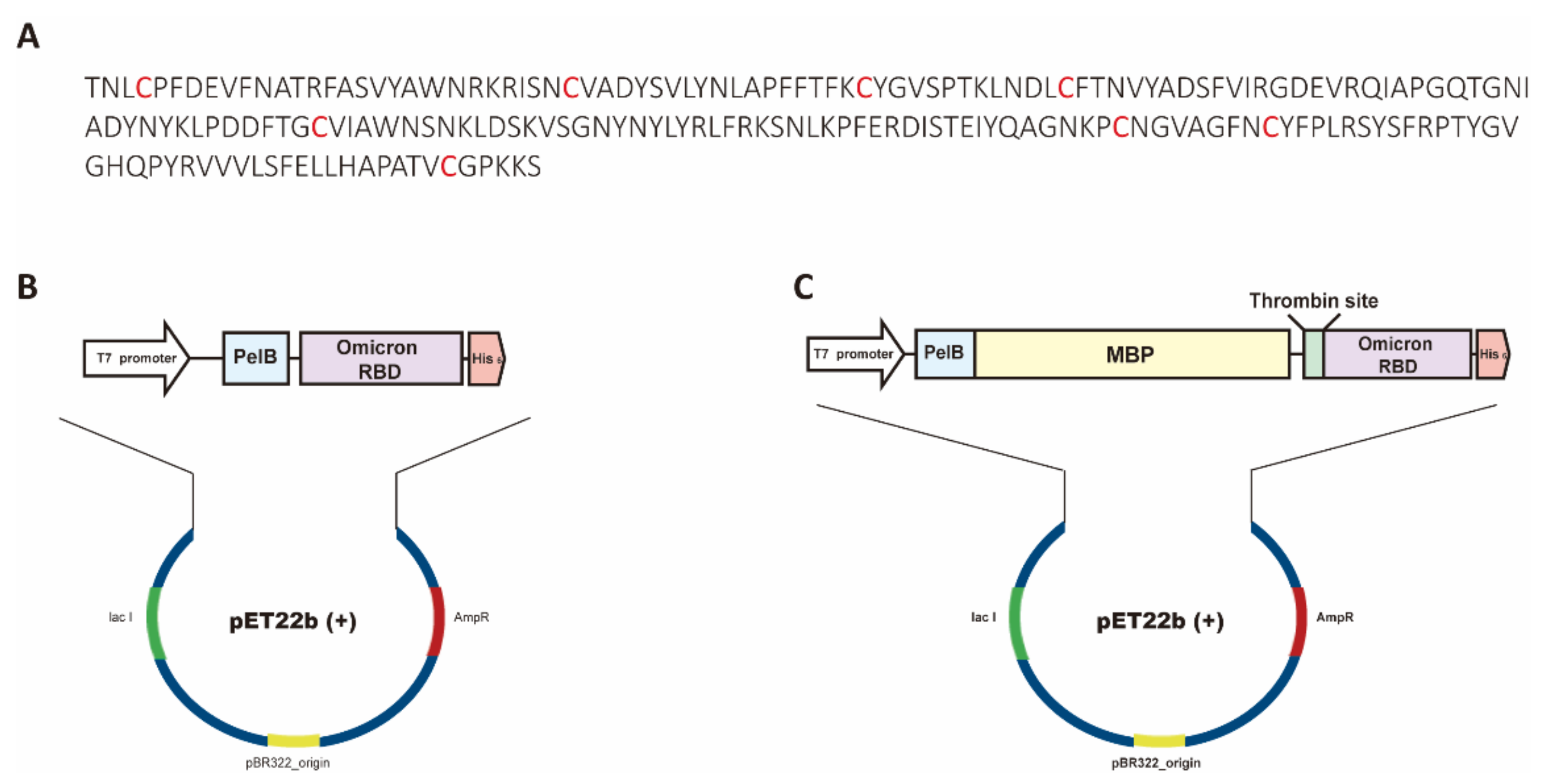
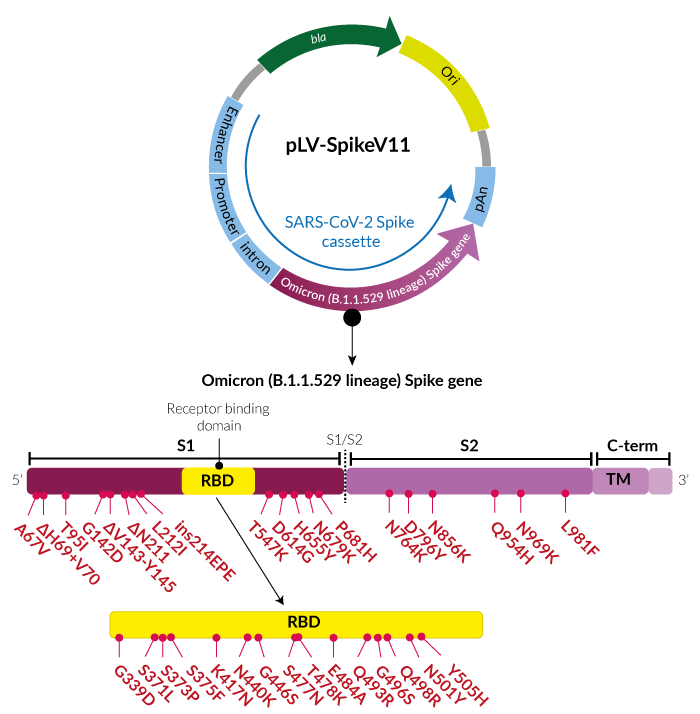
Bioengineering | Free Full-Text | Functional Expression of the Recombinant Spike Receptor Binding Domain of SARS-CoV-2 Omicron in the Periplasm of Escherichia coli
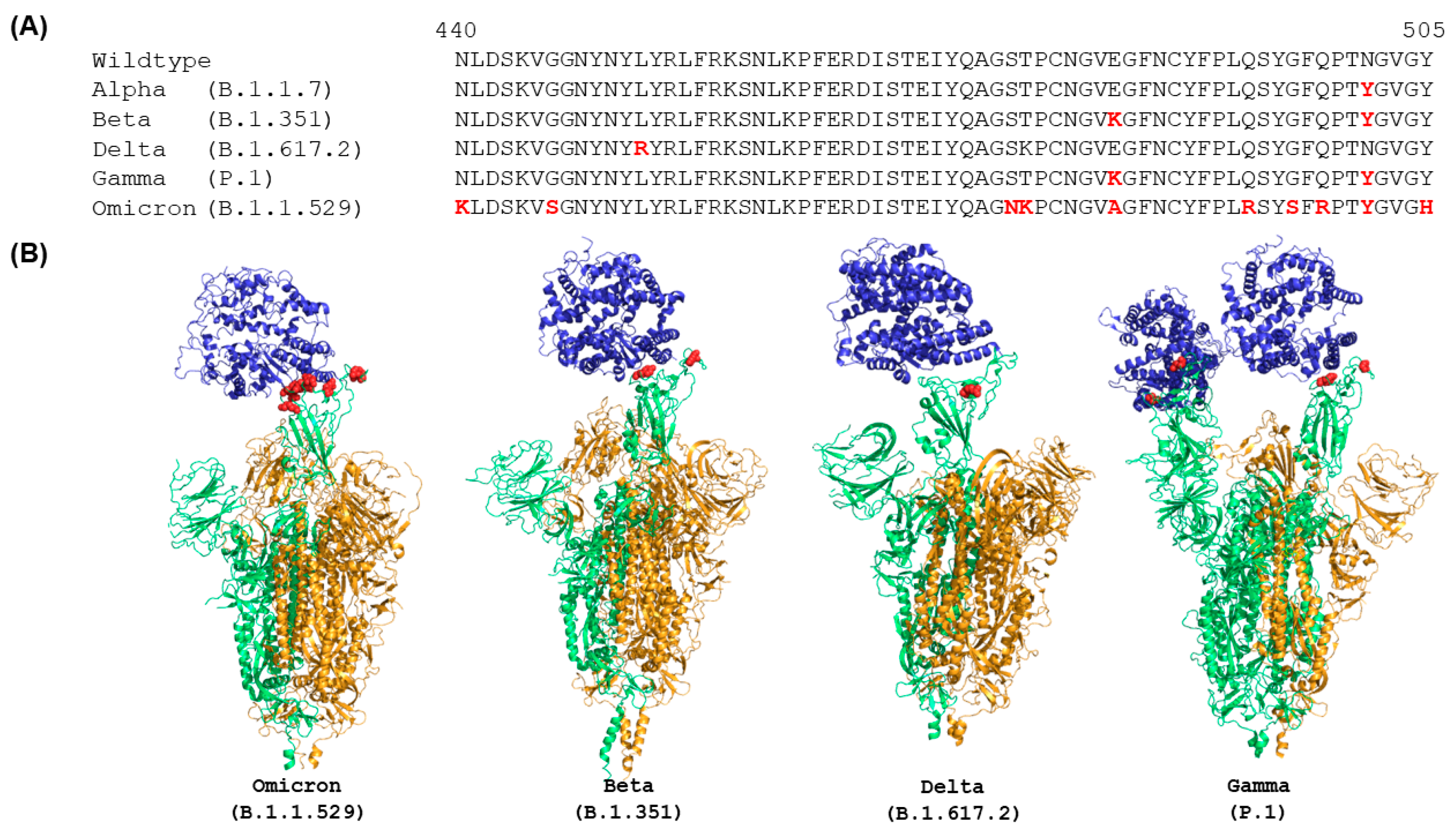
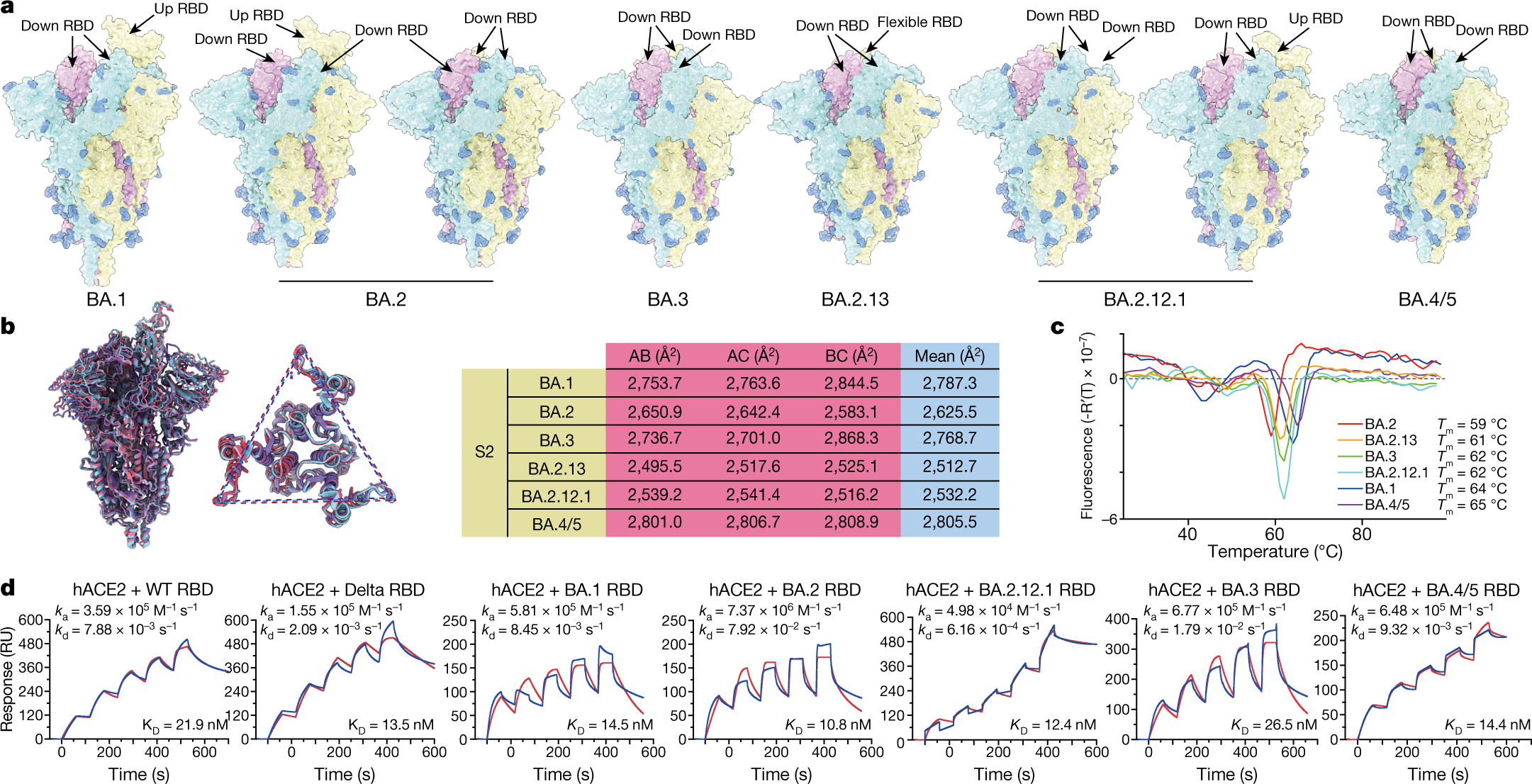
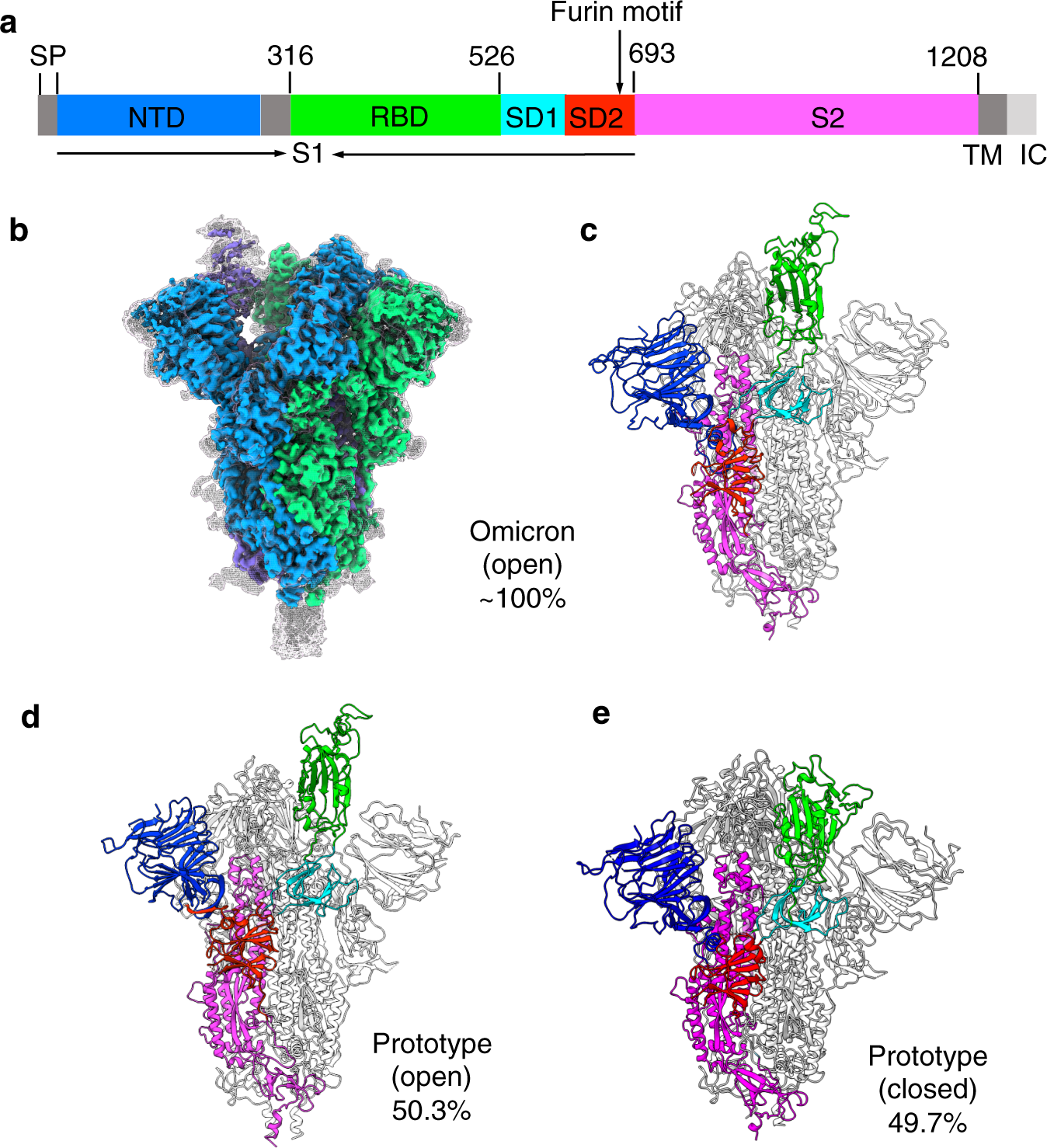
IJMS | Free Full-Text | Distinct Conformations of SARS-CoV-2 Omicron Spike Protein and Its Interaction with ACE2 and Antibody
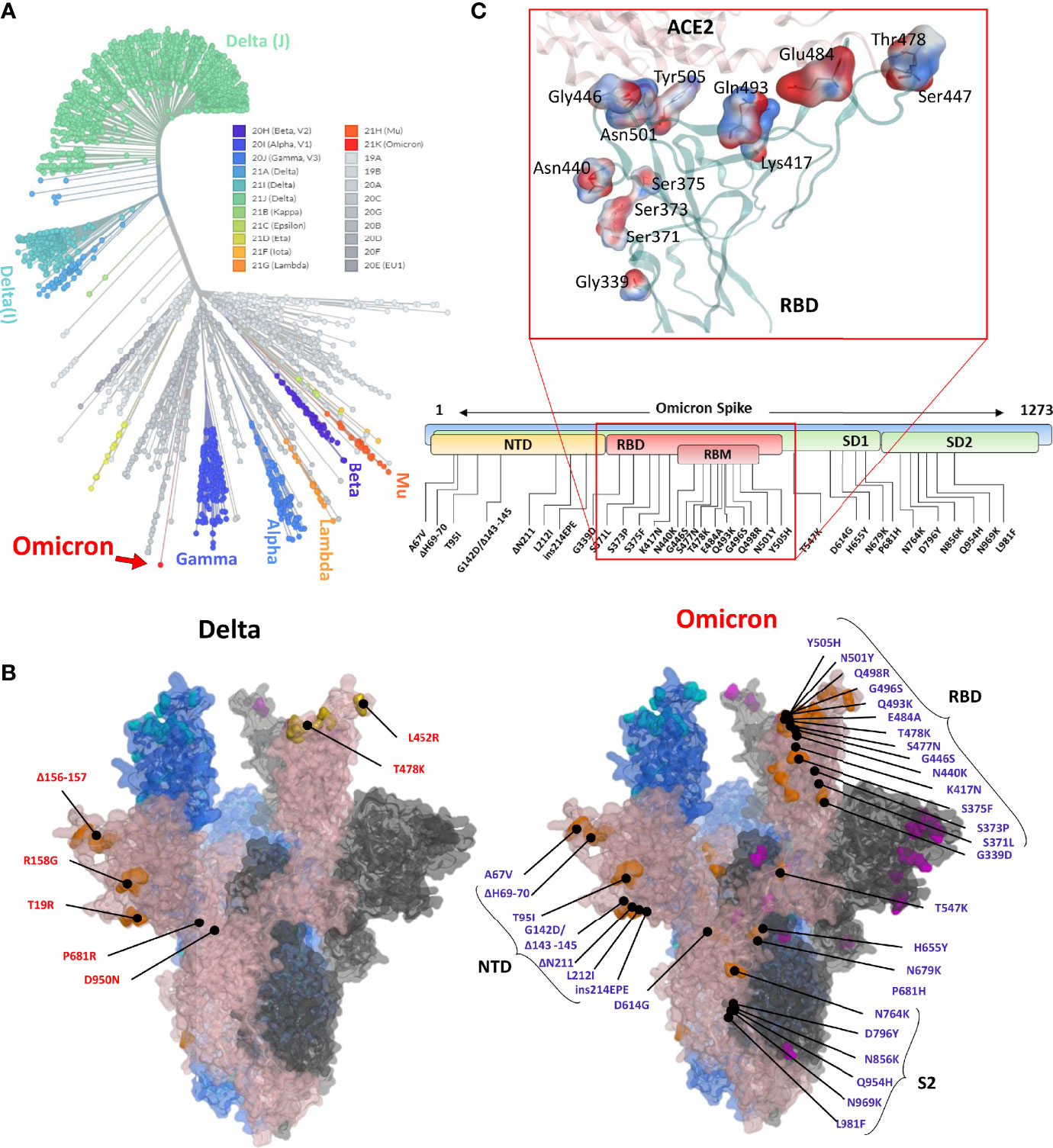
Frontiers | Omicron: A Heavily Mutated SARS-CoV-2 Variant Exhibits Stronger Binding to ACE2 and Potently Escapes Approved COVID-19 Therapeutic Antibodies
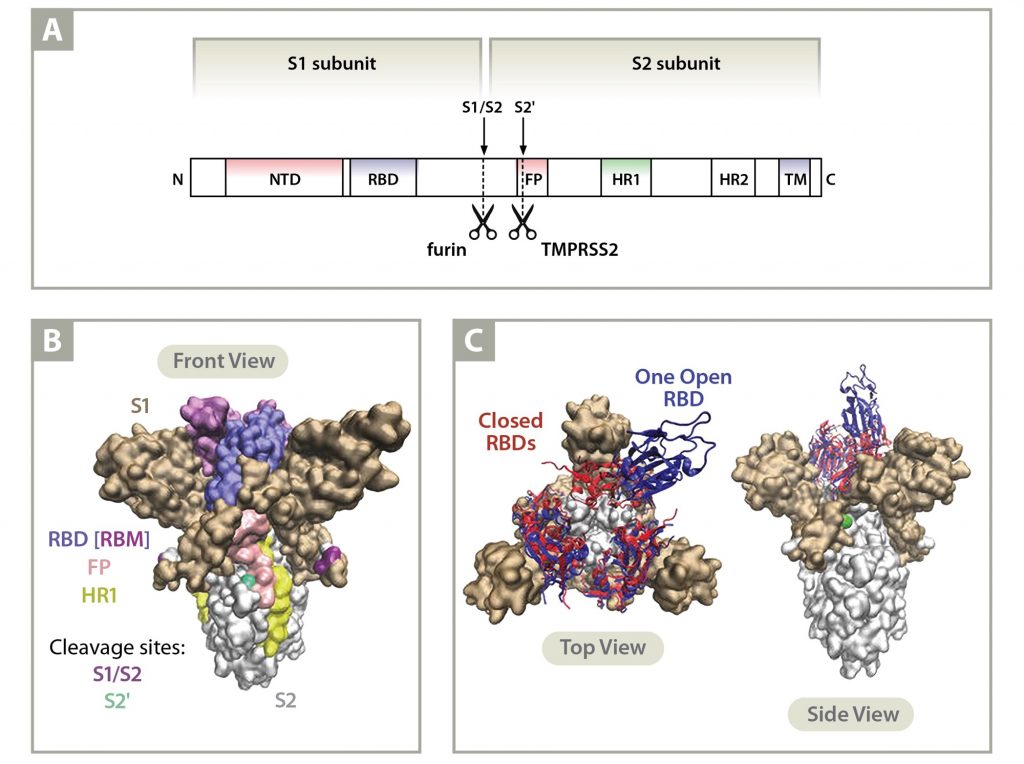
Implications of the Mutations in the Spike Protein of the Omicron Variant of Concern (VoC) of SARS-CoV-2 – Signature Science
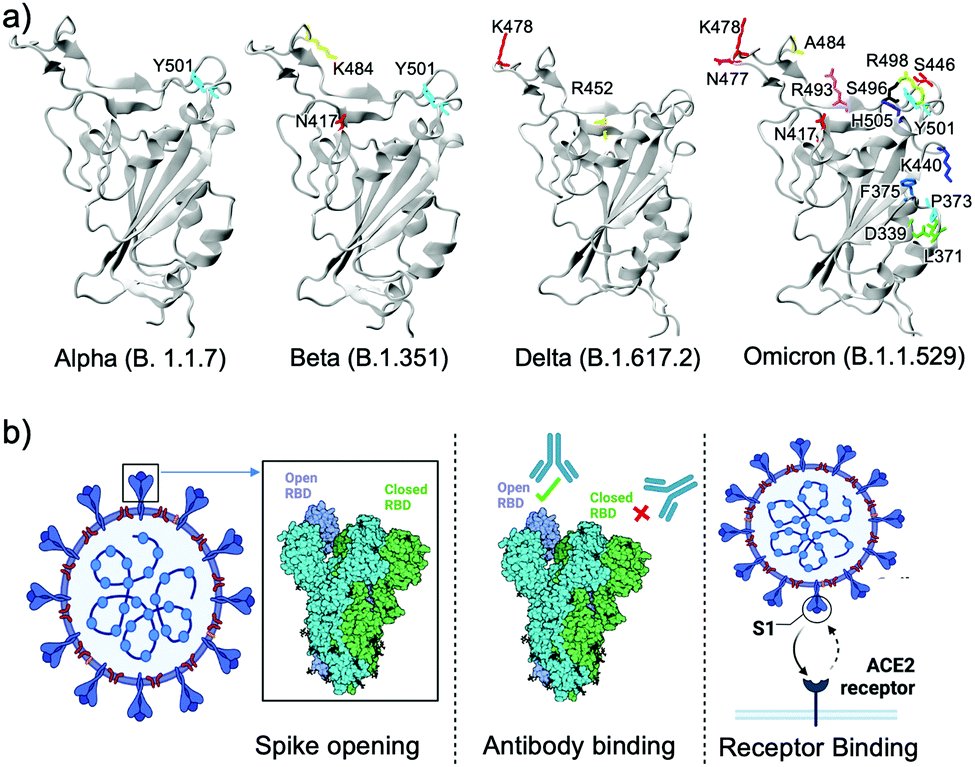
Significance of the RBD mutations in the SARS-CoV-2 omicron: from spike opening to antibody escape and cell attachment - Physical Chemistry Chemical Physics (RSC Publishing) DOI:10.1039/D2CP00169A

Mutations in the spike RBD of SARS-CoV-2 omicron variant may increase infectivity without dramatically altering the efficacy of current multi-dosage vaccinations | bioRxiv

Characterization of SARS-CoV-2 Omicron spike RBD reveals significantly decreased stability, severe evasion of neutralizing-antibody recognition but unaffected engagement by decoy ACE2 modified for enhanced RBD binding | Signal Transduction and Targeted ...
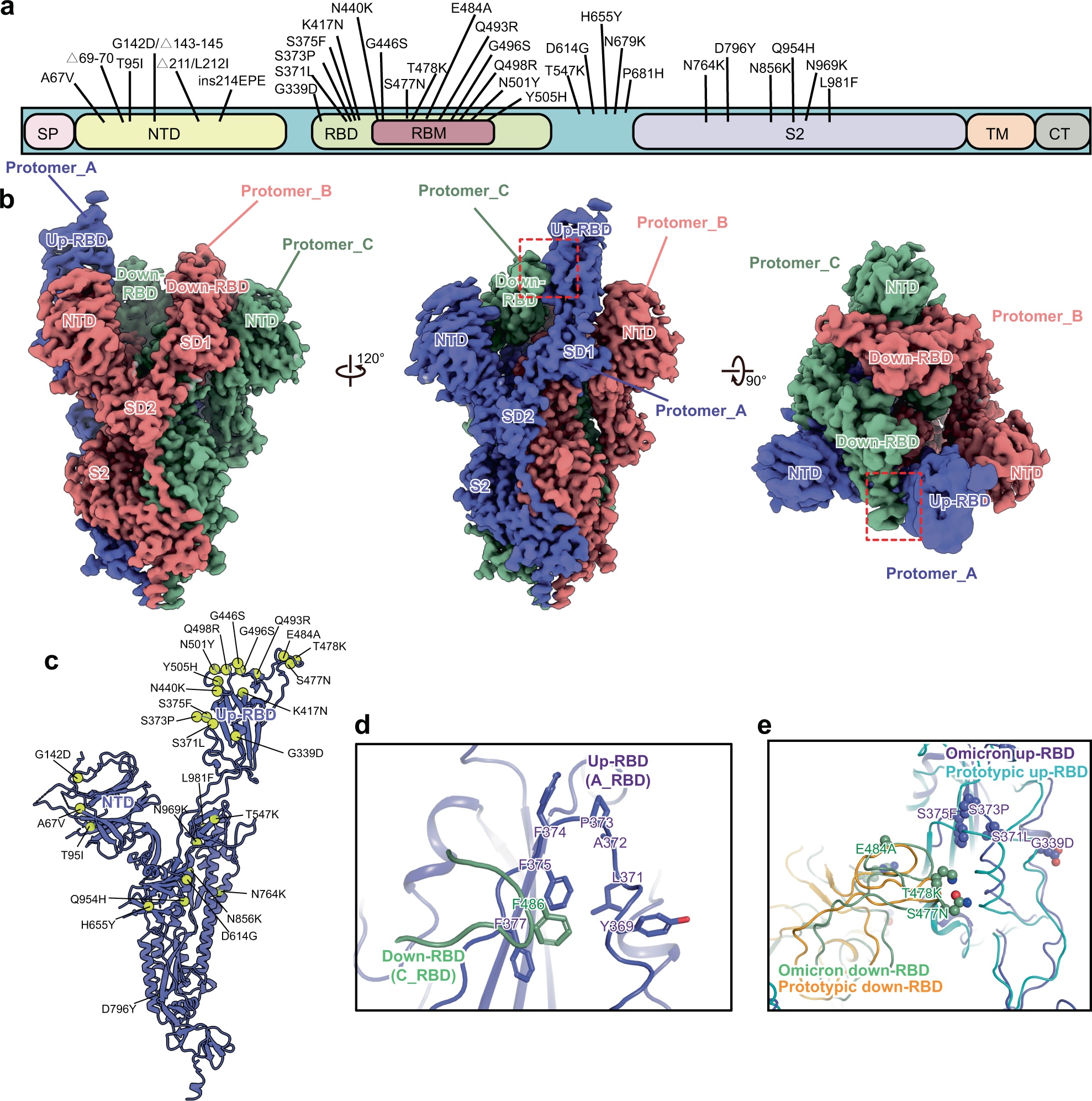
Omicron SARS-CoV-2 mutations stabilize spike up-RBD conformation and lead to a non-RBM-binding monoclonal antibody escape | Nature Communications

Sequence analysis of the emerging SARS‐CoV‐2 variant Omicron in South Africa - Wang - 2022 - Journal of Medical Virology - Wiley Online Library
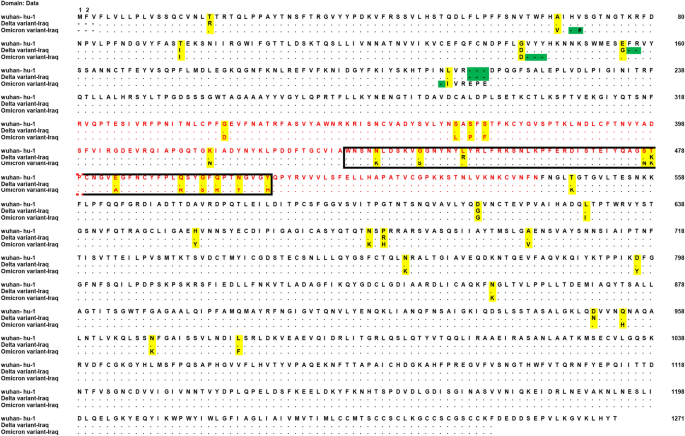
Molecular and computational analysis of spike protein of newly emerged omicron variant in comparison to the delta variant of SARS-CoV-2 in Iraq | SpringerLink