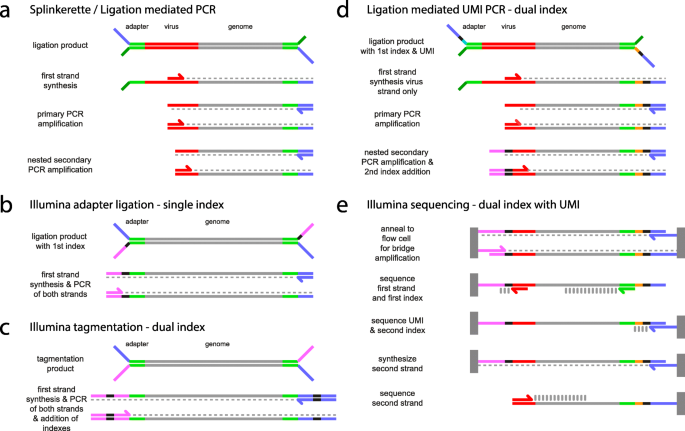
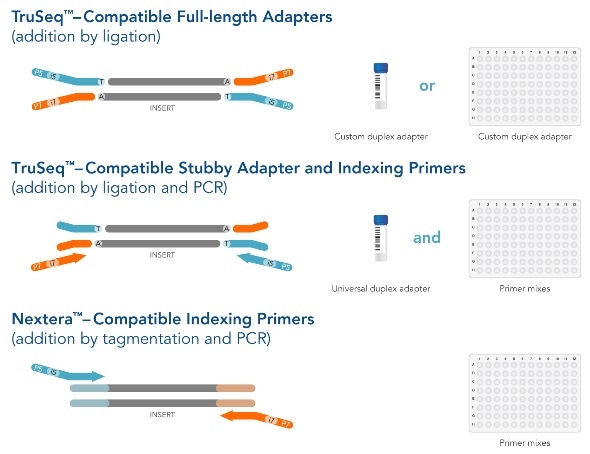
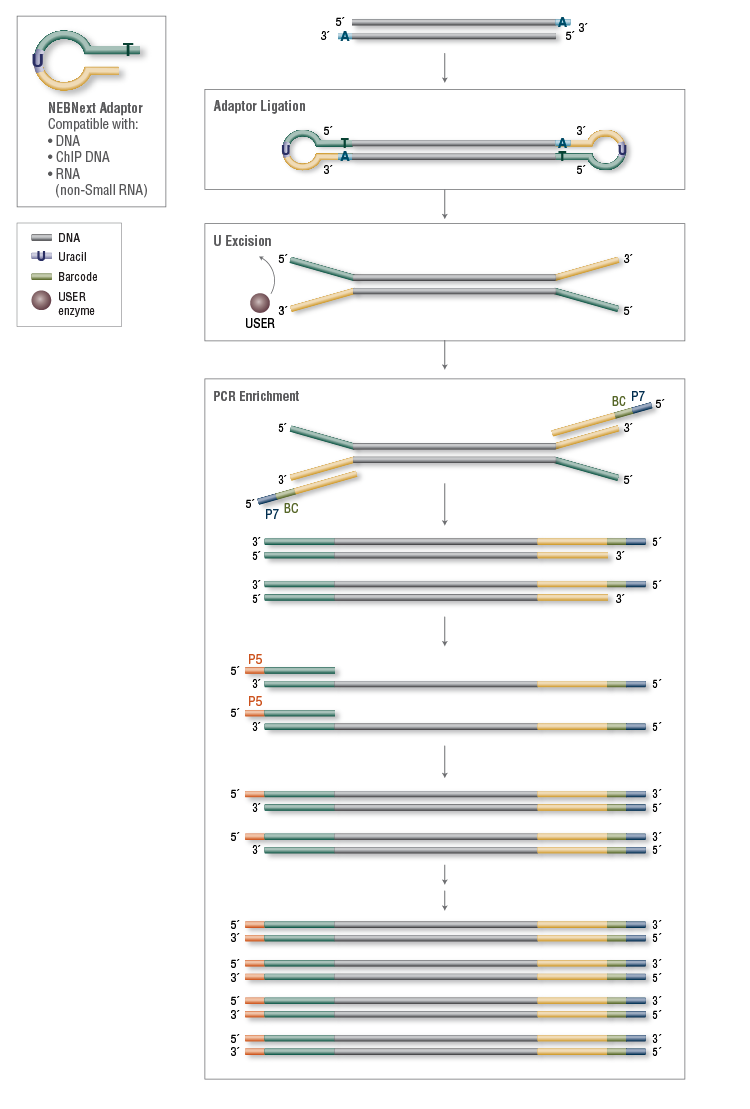
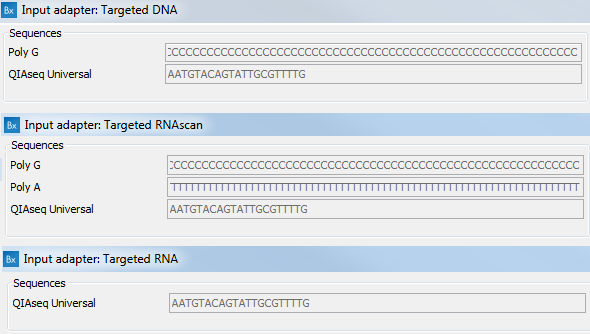
Overview of the different methods used for adapter addition to antibody... | Download Scientific Diagram
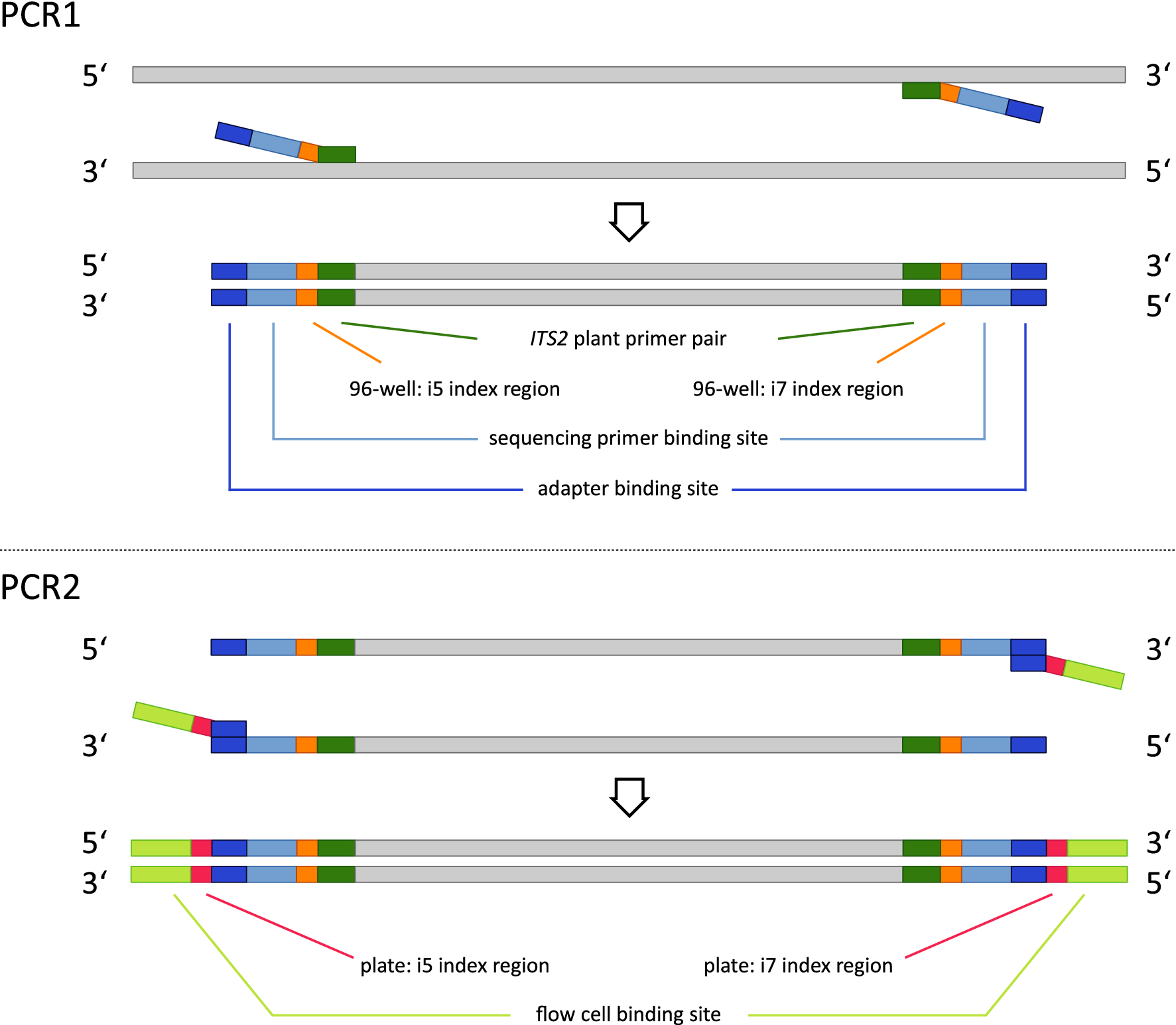
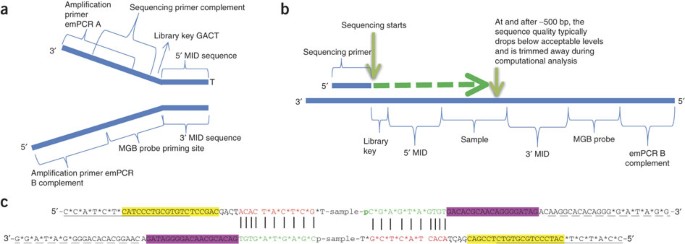
Handling of targeted amplicon sequencing data focusing on index hopping and demultiplexing using a nested metabarcoding approach in ecology | Scientific Reports

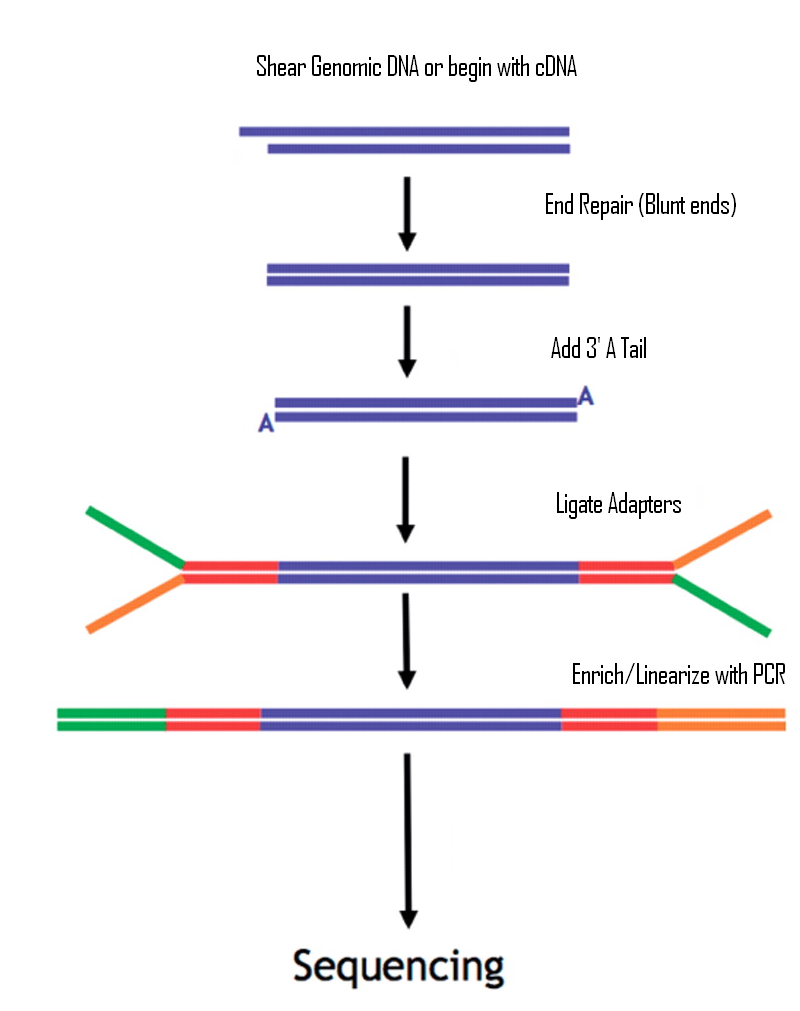
Library preparation workflow for miRNA-seq. A) The Illumina workflow... | Download Scientific Diagram

Adapter trimming: Why are adapter sequences trimmed from only the 3' ends of reads - Illumina Knowledge
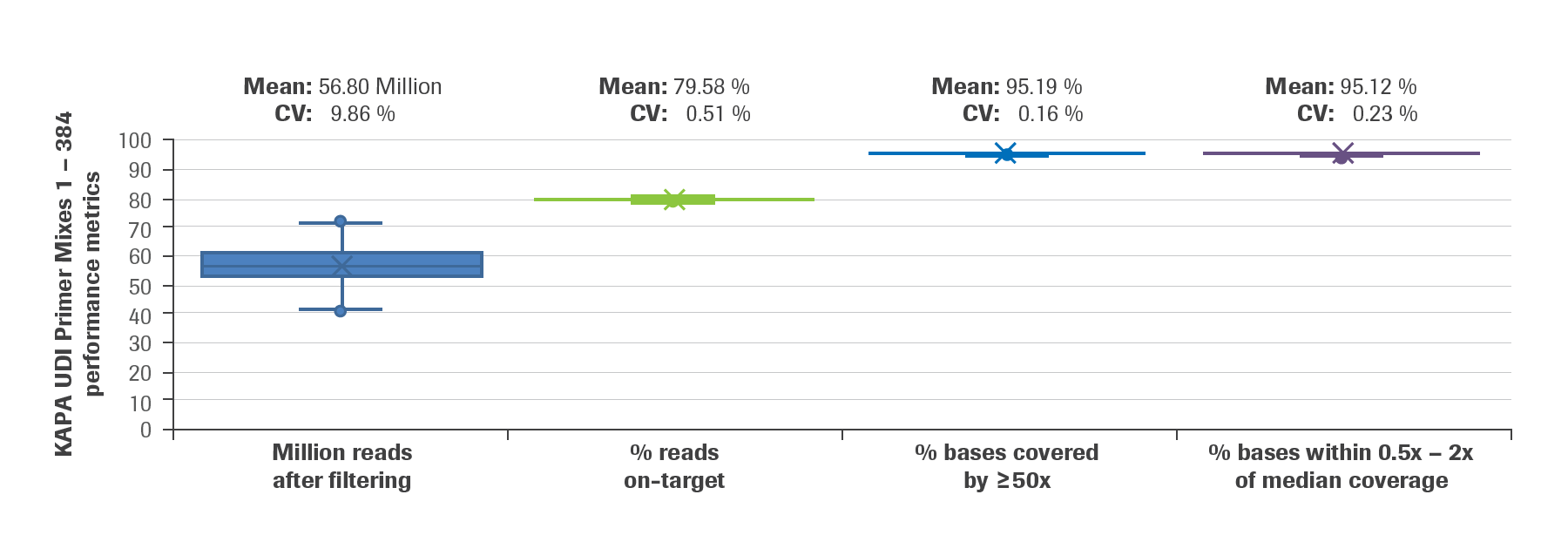
High-Throughput, Amplicon-Based Sequencing of the CREBBP Gene as a Tool to Develop a Universal Platform-Independent Assay | PLOS ONE

Traditional HPLC is Incapable of Reducing Cross Contamination of Custom Next-Generation Sequencing Adapters to Acceptable Levels
Library preparation (A) Sequence of adapters (single index) used in our... | Download Scientific Diagram

Illustrations of paired-end sequencing. (A) illustrates two strands of... | Download Scientific Diagram
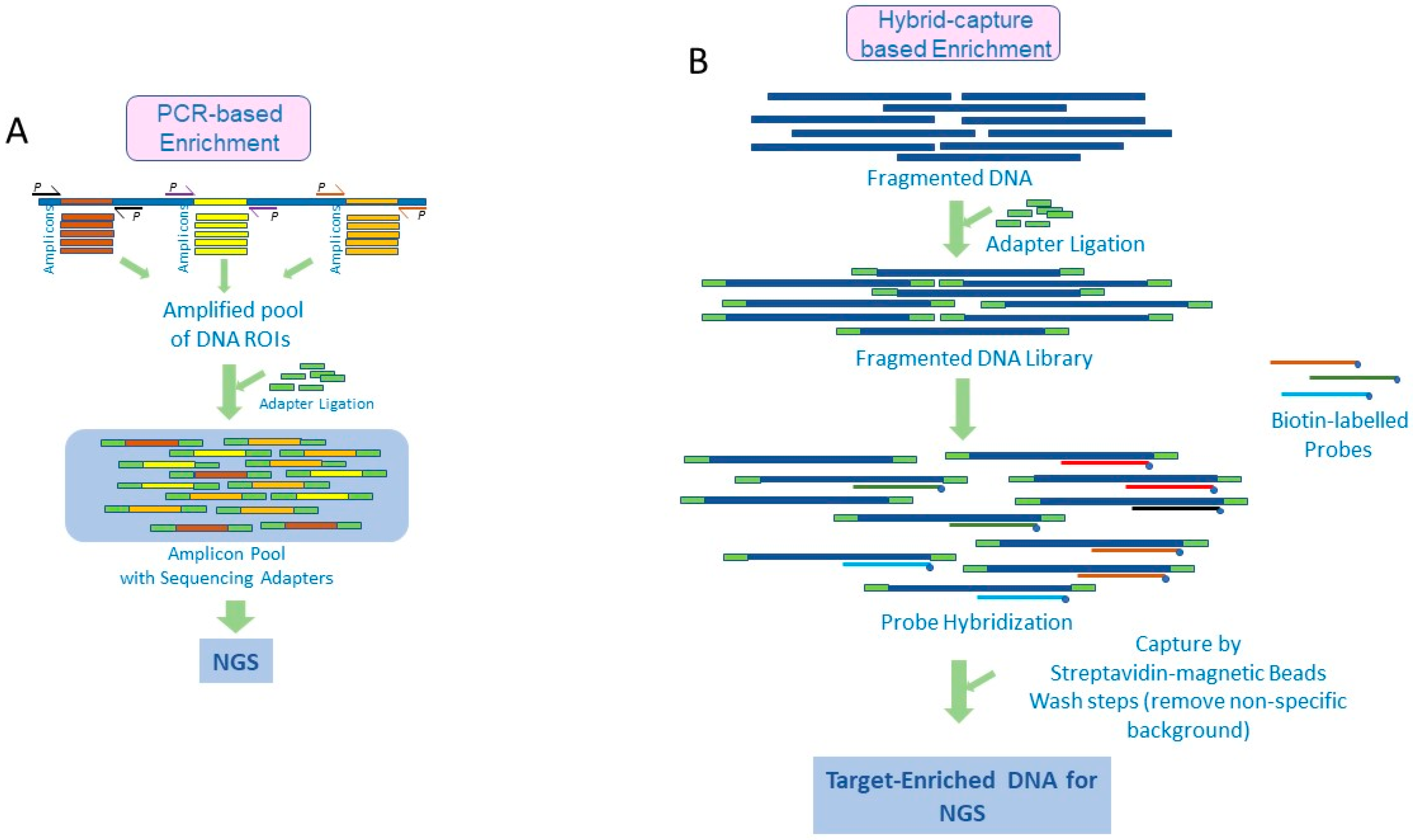
Diagnostics | Free Full-Text | Target Enrichment Approaches for Next-Generation Sequencing Applications in Oncology