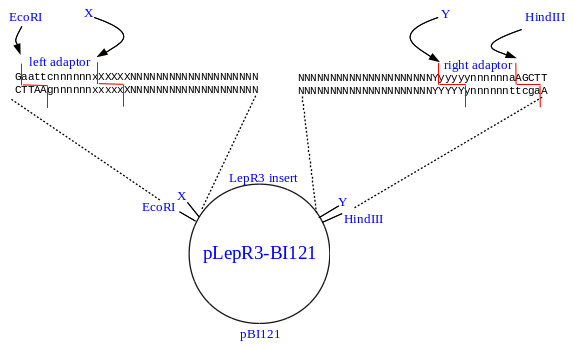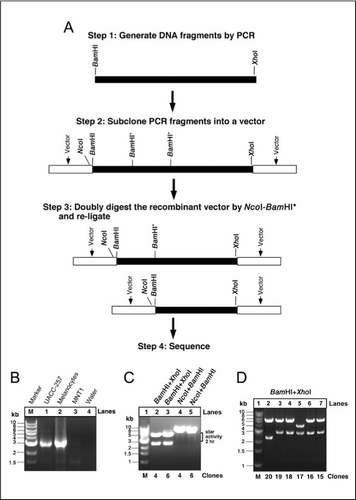
Useful tool to generate unidirectional deletion vectors by utilizing the star activity of BamHI in an NcoI-BamHI-XhoI cassette | BioTechniques

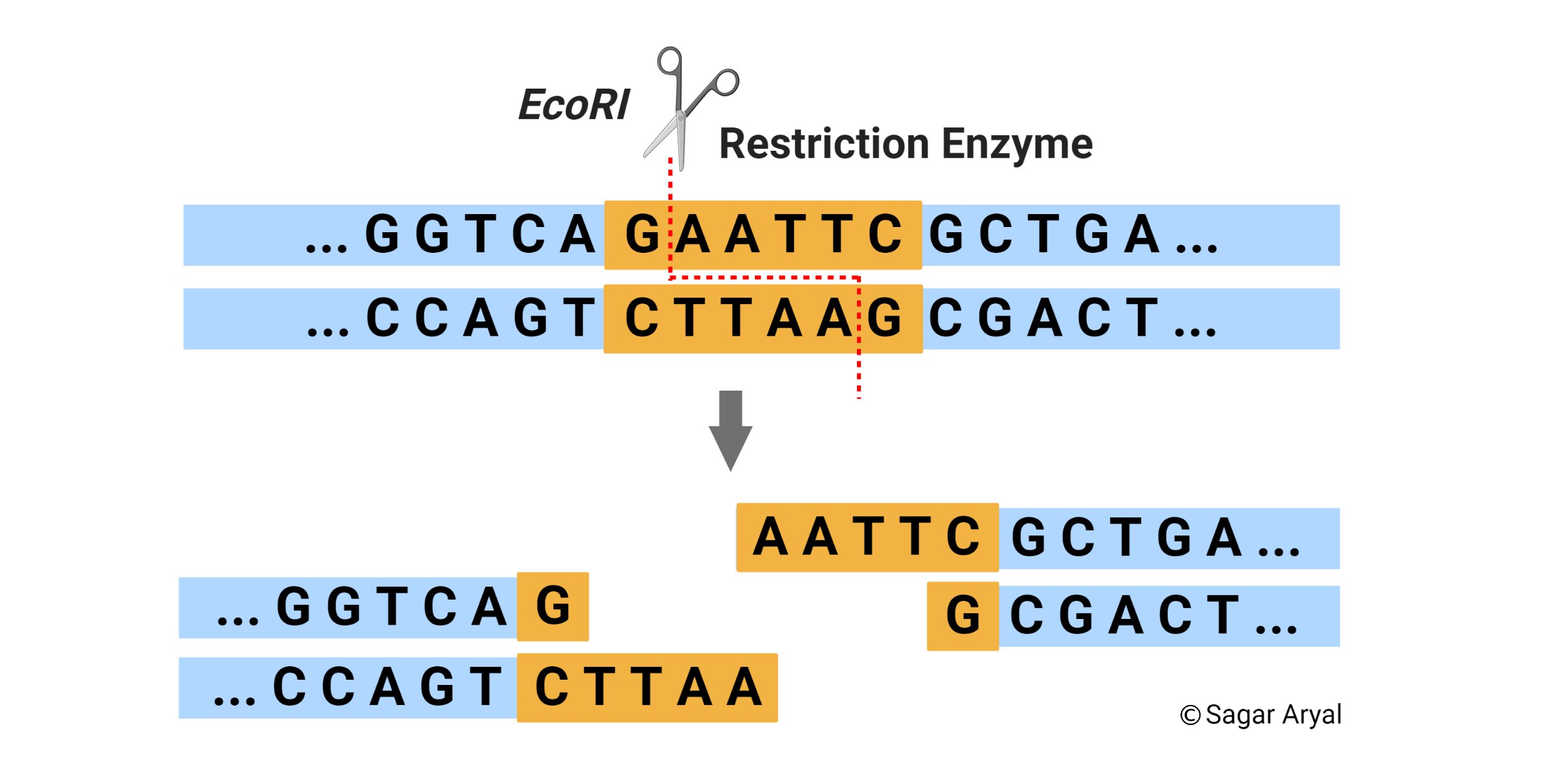
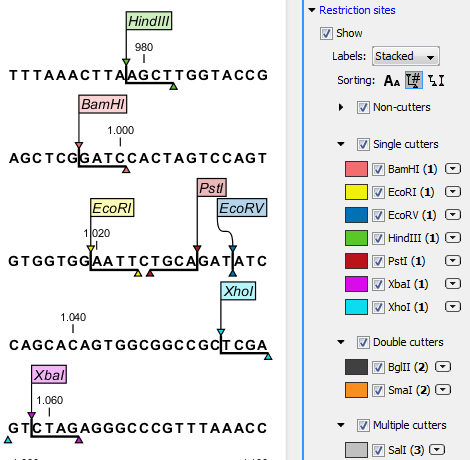
What is the Difference Between EcoRI and HindIII Restriction Enzymes | Compare the Difference Between Similar Terms

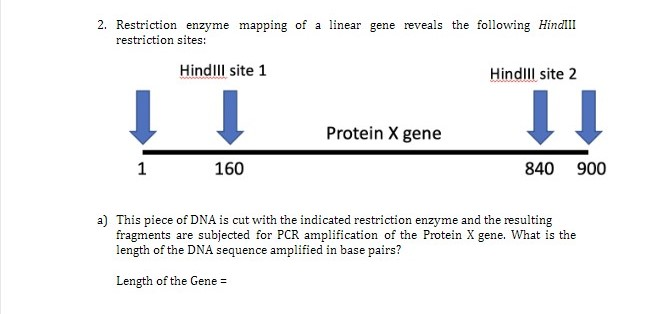
SOLVED: You have chosen to cut your DNA and plasmid with the HindIII restriction enzyme. This enzyme recognizes the sequence AAGCTT and makes a cut between the two A bases. This will