Unveiling the gene regulatory landscape in diseases through the identification of DNase I‑hypersensitive sites (Review)

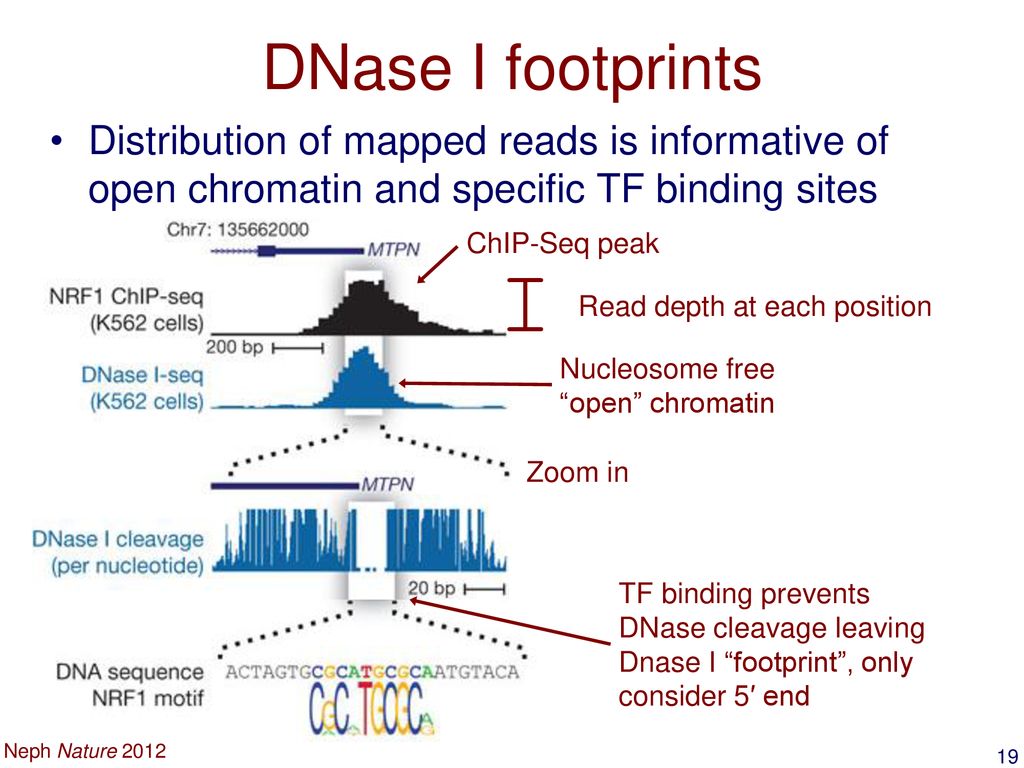
DNase Seq | DNase Footprinting | DNase1 Footprinting | DNase 1 Hypersensitive Sites Sequencing | - YouTube
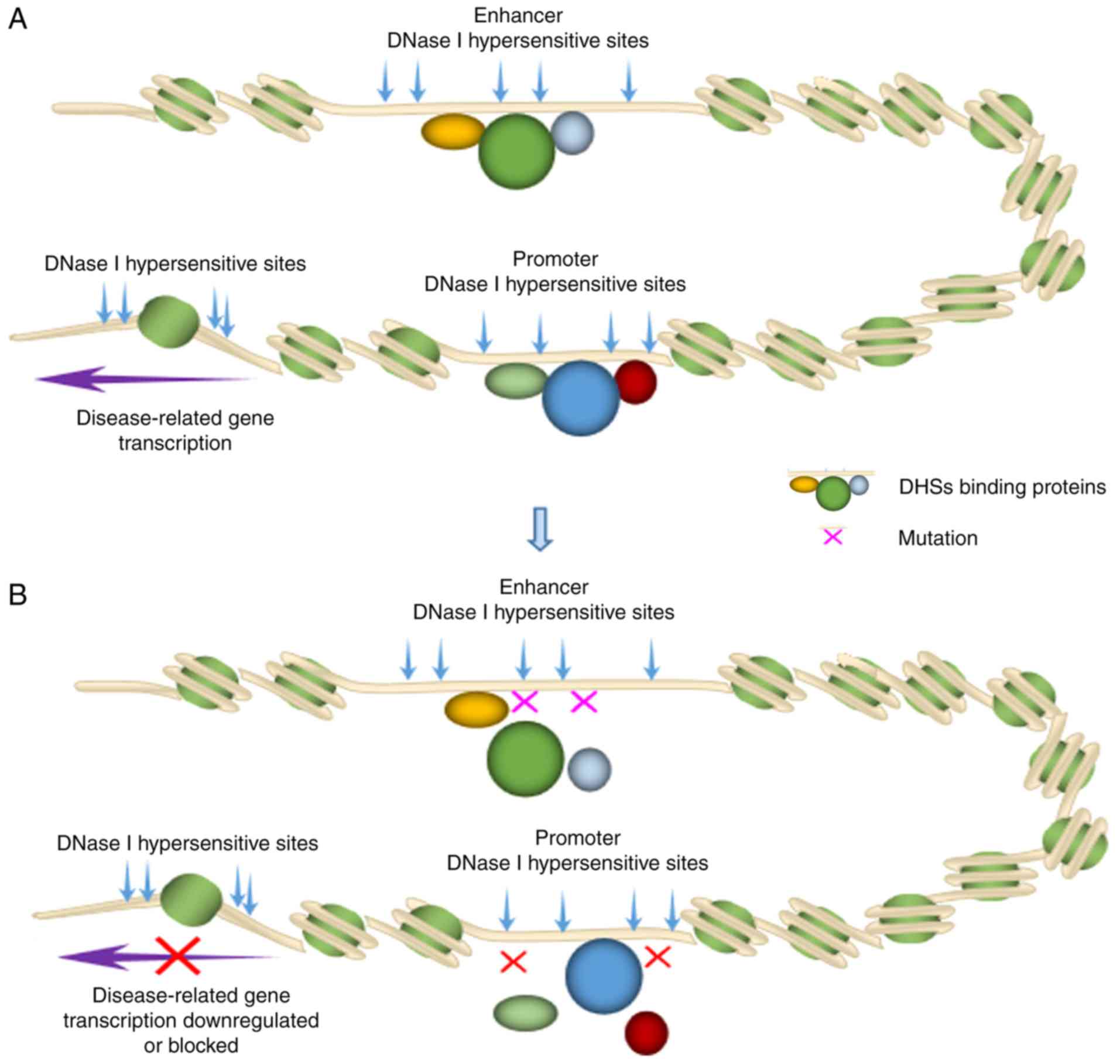
Unveiling the gene regulatory landscape in diseases through the identification of DNase I‑hypersensitive sites (Review)
DNase-seq: A High-Resolution Technique for Mapping Active Gene Regulatory Elements across the Genome from Mammalian Cells

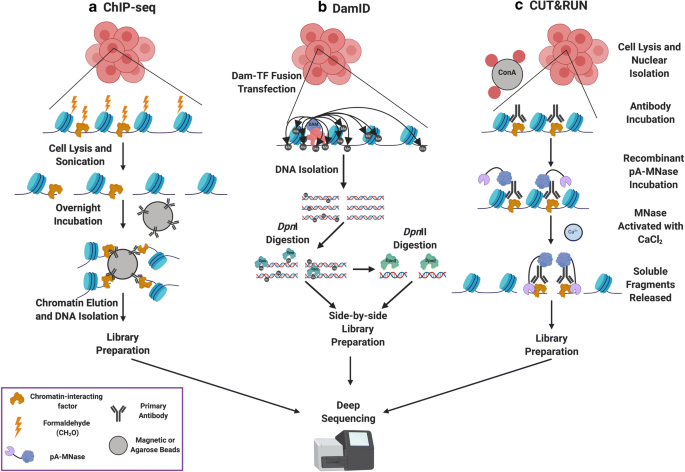
Nature Reviews Genetics on Twitter: "Principal methods for measuring chromatin accessibility: #DNAse-seq #ATAC-seq #MNase-seq #NOMe-seq Figure from our Review: Chromatin accessibility and the regulatory epigenome https://t.co/SYwpxOCv81 by @WJGreenleaf ...

Isolation of nuclei for use in genome-wide DNase hypersensitivity assays to probe chromatin structure. - Abstract - Europe PMC

DNase-seq: A High-Resolution Technique for Mapping Active Gene Regulatory Elements across the Genome from Mammalian Cells

Identifying and mitigating bias in next-generation sequencing methods for chromatin biology. - Abstract - Europe PMC
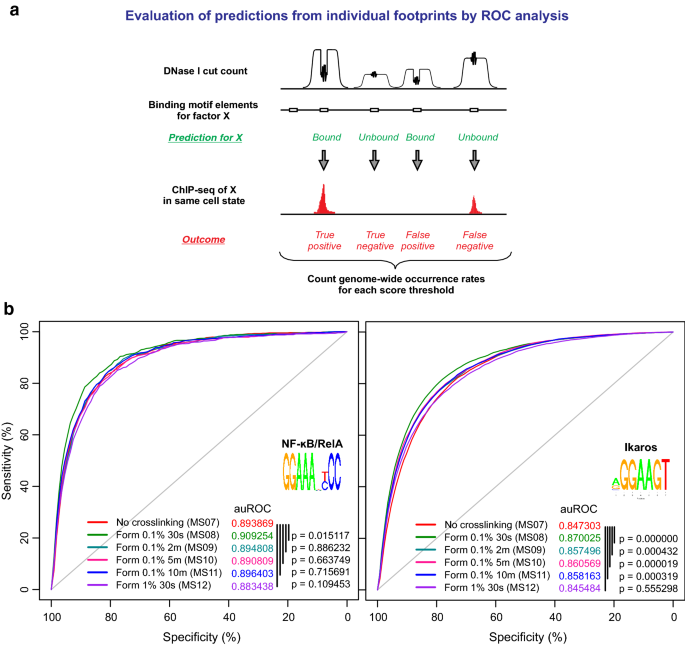
XL-DNase-seq: improved footprinting of dynamic transcription factors | Epigenetics & Chromatin | Full Text

Methods for mapping genome accessibility. A DNase-seq identifies open... | Download Scientific Diagram

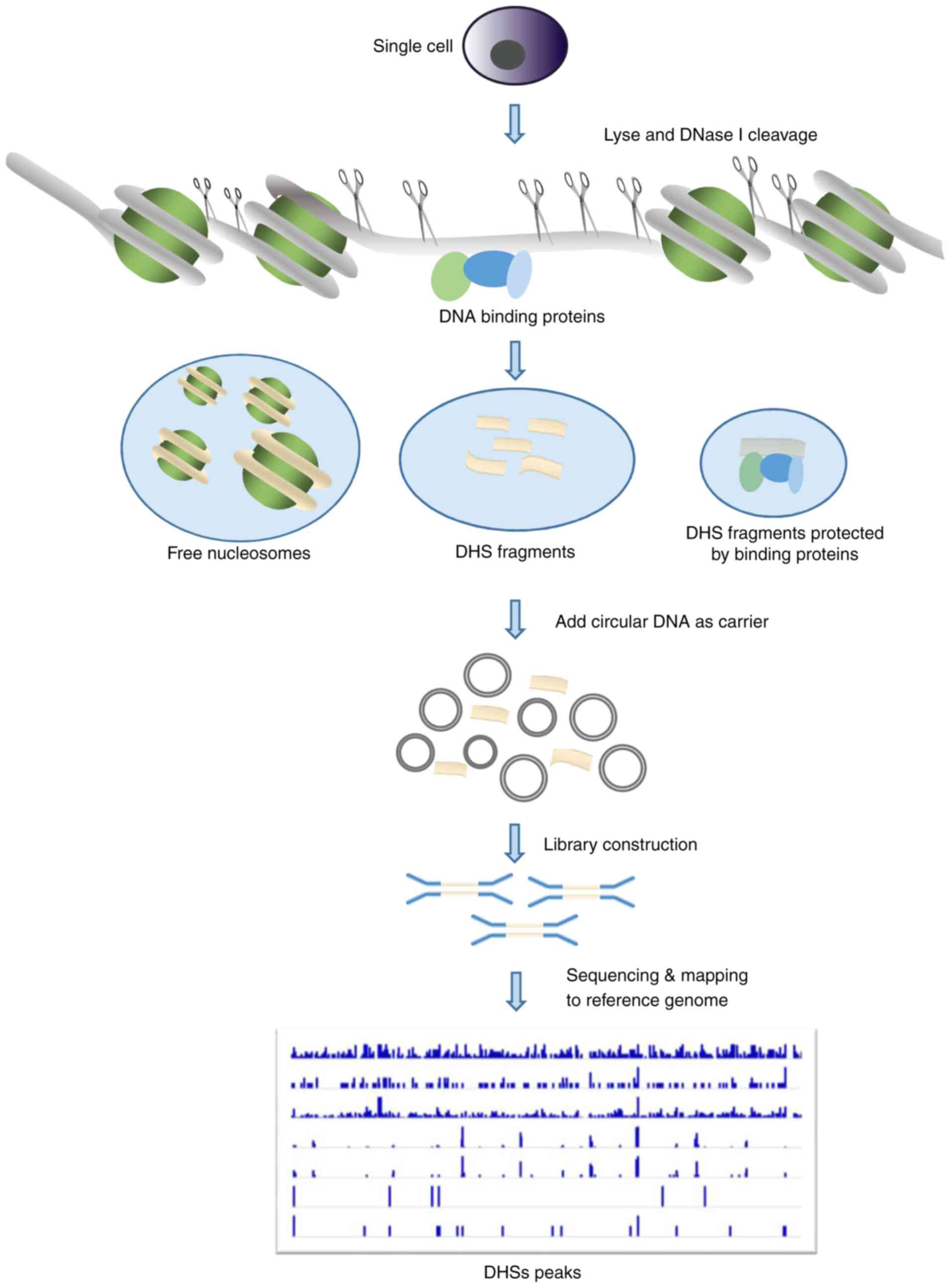
IJMS | Free Full-Text | Emerging Single-Cell Technological Approaches to Investigate Chromatin Dynamics and Centromere Regulation in Human Health and Disease
![PDF] Explicit DNase sequence bias modeling enables high-resolution transcription factor footprint detection | Semantic Scholar PDF] Explicit DNase sequence bias modeling enables high-resolution transcription factor footprint detection | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/800d7add4aada32c5e80b1e1cf6cc33319828da3/5-Figure1-1.png)